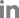|
 |
| Plant Pathol J > Volume 36(6); 2020 > Article |
|
Abstract
The type III secretion system (T3SS) is a key virulence determinant in the infection process of Pseudomonas syringae pv. tomato DC3000 (Pst DC3000). Pathogen constructs a type III apparatus to translocate effector proteins into host cells, which have various roles in pathogenesis. 4-Hydroxybenozic acid and vanillic acid were identified from root extract of Sedum middendorffianum to have inhibitory effect on promoter activity of hrpA gene encoding the structural protein of the T3SS apparatus. The phenolic acids at 2.5 mM significantly suppressed the expression of hopP1, hrpA, and hrpL in the hrp/hrc gene cluster without growth retardation of Pst DC3000. Auto-agglutination of Pst DC3000 cells, which is induced by T3SS, was impaired by the treatment of 4-hydroxybenzoic acid and vanillic acid. Additionally, 2.5 mM of each two phenolic acids attenuated disease symptoms including chlorosis surrounding bacterial specks on tomato leaves. Our results suggest that 4-hydroxybenzoic acid and vanillic acid are potential anti-virulence agents suppressing T3SS of Pst DC3000 for the control of bacterial diseases.
Antibiotics used in agriculture have been the most effective tool for bacterial disease management. However, the continuous application of antibiotics caused resistant strain development, resulting in insufficient control efficacy in fields. Streptomycin is a representative antibiotic that has been used for the control of broad-range bacterial pathogens since the early 1950s (Leben and Keitt, 1954). Streptomycin effectively managed disease development in apple and pear caused by Erwinia amylovora and Pseudomonas syringae, yet the antibiotic lost its efficacy over time because of rapid increase of streptomycin-resistant strains (Burr et al., 1988; Chiou and Jones, 1995; Spotts and Cervantes, 1995). Furthermore, possibility of horizontal transfer of resistance gene caused public concerns on triggering clinically serious problem associated with environmental and human health risks.
Anti-virulence agent modulating pathogenesis related to process such as quorum sensing (QS), biofilm formation and type III secretion system (T3SS) is currently being explored as a new strategy to control bacterial diseases in fields. Anti-virulence agent targets bacterial virulence factors without affecting the survival or propagation of bacterial pathogens. Therefore, they would have little impact on beneficial microbial flora and do not exert evolutionary pressure for the development of resistance (Dickey et al., 2017). In one of our previous studies, we isolated three resveratrol oligomers, which showed significant inhibitory effect on T3SS expression of P. syringae pv. tomato DC3000 (Pst DC3000) and effectively controlled bacterial specks and chlorosis in tomato leaves (Kang et al., 2020). Likewise, piericidin A isolated from Streptomyces xanthocidicus not only diminished the disease severity of potato tubers infected with E. carotovora subsp. atroseptica as a potential QS inhibitor, but also blocked T3SS assembly of Yersinia pseudotuberculosis with T3SS inhibitory activity (Kang et al., 2016; Morgan et al., 2017). These results demonstrate that the T3SS of pathogenic bacteria is a promising target for the development of anti-virulence agents to control plant diseases.
Pst DC3000 is a model pathogen to investigate molecular mechanism of pathogenesis in Arabidopsis and tomato plants. One of the major virulence determinants in Pst DC3000 is a special secretory apparatus called T3SS (Alfano et al., 2000). All the process including assembly of type III secretion needle complex and secretion effector proteins into host plants endows the bacterial pathogens with the ability to modulate metabolism and overcome immune response of host plants, ultimately leading to disease onset. These procedures are directed by the genes of hrp (HR and pathogenicity)/hrc (HR and conserved) gene cluster. The pathogenicity island contains regulatory genes, T3SS apparatus structural genes and effector genes encoding Avr (avirulence) and Hop (Hrp-dependent outer protein) (Alfano et al., 2000). A consensus sequence motif known as hrp box in the promoter region of these genes is recognized by HrpL, an alternative sigma factor belonging to the member of the extracytoplasmic factor family (Xiao et al., 1994). HrpL interacts with hrp box sequences which are defined as 5ŌĆ▓-GGAACCNA-N13-14CCACNNA-3ŌĆ▓ and activates the transcription of downstream open reading frame such as hrpA, avrPto, and hopP1. hrpA encodes the major structural protein of the Hrp pilus, which constructs the passage to translocate effector proteins. Hrp pilus is also observed in other gram-negative plant pathogenic bacteria including Ralstonia solanacearum, E. amylovora, and Xanthomonas campestris pv. vesicatoria as an indispensable virulence factor for disease development (Boureau et al., 2002).
Natural products have been recognized as an invaluable source for drug discovery due to their diversity in structure and properties. From 1981 to 2019, about a half of all new approved drugs was originated from natural products, including unaltered natural products, botanical drugs, natural product derivatives, synthetic drugs with pharmacophores derived from natural product and natural product mimics (Newman and Cragg, 2020). Among tremendous natural sources, phenolic compounds of plants are representative natural products possessing diverse biological activities for plant and human health (Stalikas, 2007). They have a simple common structural feature that contains aromatic ring bearing at least one hydroxyl group. As one of the wellknown phenolic acids, salicylic acid (SA; 2-hydroxybenzoic acid) is synthesized in plants to regulate flowering, heat production and defense response (Raskin, 1992). In Arabidopsis, HR is accompanied by instant accumulation of SA, which is followed by long-lasting protection against a wide range of pathogen (Ryals et al., 1996). However, when the gene involved in SA biosynthesis and biogenesis was deleted in Arabidopsis, the abilities to modulate defenses responses and accumulation of SA in response to bacterial infection were significantly impaired (Jagadeeswaran et al., 2007). In our study, a natural product library comprised of plant extracts was subject to a screening system of T3SS inhibitors against Pst DC3000. A recombinant strain of Pst DC3000 of which green fluorescent protein (GFP) expression was controlled by hrpA promoter was used to find out potential T3SS inhibitors. The extract of Sedum middendorffianum (Maximowicz) 't Hart was expected to harbor active ingredients of T3SS inhibitory activity. In this study, we identified the active ingredients from the extract of S. middendorffianum and examined their effects on cell growth, T3SS related gene transcription and disease development of Pst DC3000 to evaluate their potential use in agriculture.
Wild-type and recombinant Pst DC3000 strains were grown in KingŌĆÖ B (KB) medium or Luria-Bertani (LB) medium at 28┬░C (King et al., 1954). In order to induce T3SS, Pst DC3000 strains were grown in hrp-inducing minimal medium at 22┬░C (Huynh et al., 1989). Chloramphenicol was supplemented in the culture of the recombinant at the concentration of 25 ╬╝g/ml.
Screening of T3SS inhibitors was conducted using CA9-5 strain of which GFP expression is under the control of hrpA promoter in hrp-inducing condition (Kang et al., 2020). The method for the screening was described by Kang et al. (2020). Briefly, cell density of overnight cultured CA9-5 was adjusted with hrp-inducing minimal medium to an optical density (OD600nm) of 0.4. To screen, 98 ╬╝l of bacterial suspension was dispended into each well of a black 96-well plate and plant extracts (2 ╬╝l) were added. The plate was placed in an incubator at 22┬░C and 180 rpm. After 6 h, the fluorescence intensity of each well was detected by a HIDEX sense plate reader (HIDEX, Turku, Finland) with excitation and emission wavelength at 485 nm and 535 nm, respectively. Likewise, to identify T3SS inhibitory effect of semi-purified fractions or phenolic acids, same method describe above was used.
To purify active ingredients, 750 g of Sedum middendorffianum root was extracted twice with 70% of methanol for 12 h. The extract was concentrated in vacuo and then partitioned with same amount of hexane and ethyl acetate. The ethyl acetate fraction was concentrated to be applied onto a silica gel column chromatography. The column chromatography was performed with methanol/chloroform gradient (0-100%) to get seven fractions, each of which was tested for T3SS inhibitory effect using CA9-5 strain as described above. For further purification, sephadex LH-20 gel filtration column chromatography was conducted with the concentrated silica fraction eluted with methanol. The sephadex fraction with T3SS inhibitory effect was concentrated in vacuo and dissolved in high-performance liquid chromatography (HPLC) grade methanol. Afterwards, preparative HPLC separation with Cosmosil 5C18-MS-II column (10 ├Ś 150 mm) was carried out at a rate of 3 ml/min with a linear gradient of solvent from 20% aqueous methanol (0.04% TFA) to 35% aqueous methanol (0.04% TFA) for 20 min. Each elute peak from HPLC profile was collected and examined T3SS inhibitory effect.
Structural identification for two active ingredients was conducted with electrospray ionization mass spectrometry (ESI-MS) analysis using ESIQTRAP-3200 Mass Spectrometer (Applied Biosystems, Foster City, CA, USA). Nuclear magnetic resonance (NMR) spectra were recorded on a JEOL JNM-ECZ500R spectrometer (JEOL Ltd., Tokyo, Japan) at 500 MHz for 1H-NMR and at 125 MHz for 13C-NMR. Two-dimensional NMR such as 1H-1H correlation spectroscopy, heteronuclear single quantum correlation (HMQC), and heteronuclear multiple bond correlation (HMBC) as well as onedimensional NMR such as 1H-NMR and 13C-NMR were obtained. Based on the data, structures of two compounds were determined.
Minimum inhibitory concentration (MIC) of two phenolic acids was determined by serial dilution method in culture broth. Five serial two-fold dilutions of 4-hydroxybenzoic acid and vanillic acid were prepared to obtain final concentration range from 625 ╬╝M to 10 mM. Cells from overnight culture of four plant pathogenic bacteria (Pst DC3000, Erwinia carotovora subsp. carotovora ATCC39048, E. carotovora subsp. atroseptica BAA672, and Xanthomonas campestris pv. vesicatoria) were harvested and adjusted with LB broth to obtain a final colony forming unit (cfu) to be 5 ├Ś 105 cfu/ml. Conidial suspensions of Fusarium oxysporum f. sp. lycopersici and Colletotrichum orbiculare were prepared as described by Kim et al. (2016) to a final concentration of 105 cfu/ml. The 96-well plates containing 99 ╬╝l of bacterial or conidial suspension of plant pathogens and 1 ╬╝l of phenolic acids were incubated for 48 h at 28┬░C. MIC was defined as the lowest concentration of two phenolic acids at which no visible growth was observed. The experiment was conducted with three replicates for each treatment.
For growth curve analysis, an aliquot of Pst DC3000 cells cultured overnight in KB medium was adjusted to OD600nm of 0.2 and added into 96-well plate supplemented with 2.5 mM of 4-hydroxybenzoic acid and vanillic acid. The 96-well plate was put into shaking incubator at 28┬░C for 24 h at 200 rpm. The cell density was measured every 2 h using Optizen pop UV-vis spectrophotometer (Mecasys Co., Daejeon, Korea) for 12 h, and then it was conducted every 4 h. The measurement was performed with three replicates for each treatment.
Overnight culture of Pst DC3000 in KB medium was adjusted with hrp-inducing medium to optical density (OD600nm) of 0.4. An aliquot of Pst DC3000 cells was incubated with addition of 2.5 mM of 4-hydroxybenzoic acid and vanillic acid and 0.5% dimethyl sulfoxide (DMSO) as a negative control. After 3 h at 22┬░C, RNA from each cultured cells was isolated using a Nucleospin RNA kit (Macherey-Nagel, D├╝ren, Germany) according to the manufacturerŌĆÖs protocol. The quantity of extracted RNA was determined by Nanodrop ND-1000 spectrophotometer (NanoDrop Technologies Inc., West Lothian, UK), and then 500 ng of RNA was reverse transcribed using iScript cDNA synthesis kit (Bio-Rad, Hercules, CA, USA). For quantitative PCR, 4 ╬╝l of cDNA template, 1 ╬╝l of 10 pmol of primer set (Kang et al., 2020), 5 ╬╝l of nuclease free H2O and 10 ╬╝l of MG 2├Ś qPCR MasterMix SYBR Green (MG Med, Seoul, Korea) were transferred into PCR 96-well plates. The plate was placed into Bio-rad CFX96 (Bio-Rad) for PCR amplification. Based on the expression of internal control genes (recA), the relative expression of gyrA, hopP1, hrpA, and hrpL was calculated. The experiment was independently repeated three times with three replicates for each treatment.
To observe auto-agglutination of Pst DC3000, mutant strain (DC-N7) was constructed in our previous study (Kang et al., 2020). Strain DC-N7 contains GFP-expressing vector under the control of constitutive promoter (pnptII) to visualize bacterial autoagglutination on microscopic observation. A single colony of strain DC-N7 grown on LB agar plate overnight was inoculated into LB medium and incubated for 24 h at 28┬░C. DC-N7 cells were harvested and re-suspended in hrpinducing medium with optical density (OD600nm) of 0.5. An aliquot of the culture was incubated in 96-well plate with addition of 2.5 mM of 4-hydroxybenzoic acid and vanillic acid and 0.5% DMSO as negative control at 22┬░C for 48 h at 90 rpm. Subsequently, 20 ╬╝l of the culture was placed onto slide glass to observe auto-agglutination under a Zeiss Axio Imager M1 microscope (Carl Zeiss, Gottingen, Germany).
Tomato seeds (Solanum lycopersicum cv. Gwangbok) were immersed in water to germinate and placed in a dark room at 28┬░C for 5 days. The germinated seeds were transferred into a 32-well plastic tray (58 mm ├Ś 58 mm ├Ś 63 mm) filled with sterilized Baroker soil (Seoul Bio, Eumseong, Korea). After 3 weeks, tomato seedlings were randomly selected to examine the effect of two phenolic acids on disease development. 2.5 mM of two phenolic acids were respectively mixed with bacterial suspension of Pst DC3000 which was prepared by the same method of Kang et al. (2020). Syringe-infiltration was conducted to infect tomato leaves and then, they were placed in a humidified chamber for 48 h at 26┬░C. Disease symptoms on the tomato leaves were observed for 1 week. The experiment was independently conducted three times with three replicates of each trial.
Cfu of Pst DC3000 in the tomato leaves treated with phenolic acid mixture was measured as follows. Phenolic acid mixture was prepared from the active fractions obtained from sephadex column chromatography of S. middendorffianum extract performed as described above. The phenolic acid mixture (17.8 ╬╝g of 4-hydroxybenzoic acid and 7.2 ╬╝g of vanillic acid in 25 ╬╝g of the mixture) was mixed with 200 ╬╝l of Pst DC3000 cell suspension in 10 mM of sucrose (5 ├Ś 103 cfu/ml). The bacterial suspension (200 ╬╝l) mixed with or without phenolic acid mixture was injected into each tomato leaf. After 7 days, the inoculated lesions were cut into 1 cm ├Ś 1 cm pieces and then homogenized in 1 ml of sterilized water. The suspension was further diluted and plated onto KB medium to measure cfu. The plates were incubated in 28┬░C for 2 days. Each treatment had three replicates.
For statistical analysis of experimental data, Statistical Analysis System software (SAS version 9.4, SAS Institute Inc., Cary, NC, USA) was used. The significant differences in means of the data were determined by One-way analysis of variance (ANOVA) with using the least significant difference test at P < 0.05.
Through screening of natural product library, the extracts of Sedum species including Sedum middendorffianum and S. kamtschaticum were selected to potentially harbor inhibitors of GFP expression under the control of hrpA promotor. The root and shoot extracts of S. middendorffianum and S. kamtschaticum were examined to determine which parts of the plants possess better inhibitory activity. Both parts of S. middendorffianum showed inhibitory effect of GFP expression (Fig. 1). The root extract of S. middendorffianum displayed the most predominant reduction in fluorescence intensity (Fig. 1), which suppressed the fluorescence intensity to 11% of the control. Therefore, we focused on the identification of active ingredients from the extract of S. middendorffianum, because it had better inhibitory effect on GFP expression.
The root of S. middendorffianum was extracted with 70% aqueous methanol and partitioned with hexane and ethyl acetate. Among the fractions, the ethyl acetate extract possessed potential T3SS inhibitors. After silica column chromatography of the extract, active fractions 51 to 76 were re-fractionated by sephadex LH-20 column chromatography. Total fractions were grouped into 8 pooled fractions according to their thin-layer chromatography patterns. The second pooled fraction showed the most inhibitory activity on fluorescence intensity of CA9-5 strain. Further purification was conducted using preparative HPLC. The eluates of two peaks at 13.2 min and 13.6 min retention time reduced fluorescence intensity which were named as compound A and compound B (Supplementary Fig. 1).
Compound A was obtained as a white powder. Mass spectra of compound A recorded with a deprotonated molecular peak at 137.1. The 1H-NMR spectra of compound A revealed that compound A has only two signals of aromatic methine protons. In the 13C-NMR spectrum, 5 signals including one carbonyl, one oxygenated sp2 quaternary, two sp2 methine and one sp2 quaternary carbon were detected. By means of HMQC spectra, two protons were respectively associated with each carbon. HMBC correlations were observed between methine proton at 6.81 ppm and sp2 quaternary carbon at 122.7 ppm and sp2 methine carbon at 116.0 ppm, and between methine proton at 7.87 ppm and carboxyl and oxygenated sp2 quaternary carbon at 170.1 ppm and 163.3 ppm. Based on these data, compound A was assigned as 4-hydroxybenzoic acid (Fig. 2A).
Compound B was obtained as brown powder from the eluate of preparative HPLC. Compound B gave deprotonated ion at 166.8 in ESI-MS. The 1H-NMR spectra of compound B showed three aromatic methine protons at 7.55, 7.54, and 6.83 ppm and one methoxy proton at 3.88 ppm. The 13C-NMR and HMBC correlations recorded 8 carbon signals including carboxyl carbon, two oxygenated sp2 quaternary carbons, three sp2 methine carbons, sp2 quaternary carbon and methoxyl carbon. The HMBC spectra recorded the long-range correlations between the methine proton at 6.83 ppm and oxygenated sp2 quaternary carbon and sp2 quaternary carbon at 148.7 ppm and 123.1 ppm, and between methine protons at 7.55 and 7.54 ppm and carboxyl carbon and oxygenated sp2 quaternary carbon at 170.0 and 152.6 ppm. Methine proton at 7.55 ppm and methoxy proton at 3.88 ppm showed long-range correlation with oxygenated sp2 quaternary carbon at 148.7 ppm. These NMR spectra indicated that compound B was vanillic acid (Fig. 2B).
Antimicrobial activity of 4-hydroxybenzoic acid and vanillic acid was evaluated against various plant pathogens by MIC assay. Two-fold serial dilutions of two phenolic acids were tested for antimicrobial activity in 96-well plate. At 10 mM of 4-hydroxybenzoic acid, antibacterial activities against plant pathogenic bacteria used in this assay were observed, while the growth of two plant pathogenic fungi was not affected by 4-hydroxybenzoic acid up to this concentration (Table 1). In case of vanillic acid, MICs against Pst DC3000, E. carotovora subsp. carotovora ATCC39048, X. campestris pv. vesicatoria, F. oxysporum f. sp. lycopersici, and C. orbiculare were determined to be 10 mM while the growth of E. carotovora subsp. atroseptica BAA672 was inhibited at 5 mM (Table 1).
To evaluate the effect of 4-hydroxybenzoic acid and vanillic acid at a sub-lethal dose (1/4 MIC, 2.5 mM) on bacterial growth, growth curve analysis of Pst DC3000 was conducted (Fig. 3). Phenolic acids did not impaired growth of Pst DC3000 compared to the growth of untreated control throughout all phases. The result indicates that the growth of Pst DC3000 was not affected at 1/4 MIC of 4-hydroxybenzoic acid and vanillic acid. 1/4 MIC was the concentration displaying significant reduction of fluorescence in strain CA9-5 (Fig. 4).
To quantify the inhibitory effect of two phenolic acids on hrpA promoter activity, fluorescence intensity of strain CA9-5 was measured at varying dosages of 4-hydroxybenzoic acid and vanillic acid. 4-hydroxybenzoic acid did not reduce the fluorescence intensity until the concentration of 1.25 mM (Fig. 4A). However, the fluorescence intensity at the concentration of 2.5 mM significantly decreased to 43% of non-treated control. The intensity further decreased by 5 mM of 4-hydroxybenzoic acid treatment to 4% of the control level. In the case of vanillic acid, significant reduction was observed at the concentration of 1.25 mM recording the fluorescence intensity to be 72% of non-treated control (Fig. 4B). The fluorescence intensity was further reduced to 40% and 6% of the control at the concentration of 2.5 mM and 5 mM, respectively.
The relative transcriptional levels of hopP1 (harpin gene: secreted type III effector), hrpA (Hrp pilus gene: structural gene) and hrpL (regulatory gene) were examined by qRT-PCR to investigate the inhibitory effect of 4-hydroxybenzoic acid and vanillic acid on the expression of T3SS-related genes of Pst DC3000. In the presence of 4-hydroxybenzoic acid, the transcriptional levels of hopP1, hrpA, and hrpL were significantly below than that of non-treated control (Fig. 5). Especially, the normalized gene expression value of hrpA gene was observed below 0.01. Vanillic acid also showed the similar results in the expression levels of the three genes (Fig. 5). The expression of housekeeping gene gyrA which is responsible for DNA breakage and reunion was not affected by the treatment of two phenolic acids (Fig. 5). These results suggested that 4-hydroxybenzoic acid and vanillic acid modulate the expression of T3SS related genes in Pst DC3000.
In previous study, it was proved that T3SS inhibitors disturbed T3SS-mediated agglutination of Pst DC3000 under hrp-inducing condition (Kang et al., 2020). In this study, using DC-N7 strain expressing reporter gfp gene fused to constitutive promoter (PnptII), auto-agglutination of DC-N7 strain was visualized by fluorescence microscope in the presence of 4-hydroxybenzoic acid and vanillic acid. High number of aggregates were observed in preparation of control (Fig. 6A). However, few aggregates were detected in the preparation of bacteria treated 2.5 mM of 4-hydroxybenzoic acid and vanillic acid (Fig. 6B and C). These results indicated that auto-agglutination activity of Pst DC3000 was affected by two phenolic acids which have T3SS inhibitory effect.
Tomato plant is a host of Pst DC3000 causing bacterial speck and chlorosis. 4-Hydroxybenzoic acid and vanillic acid displaying T3SS inhibition activity were examined that they could suppress disease development in tomato plants. On Pst DC3000-infected leaves, apparent disease symptoms including chlorosis surrounding specks were observed (Fig. 7B). While the initial infection site was the left side of the leaf, weak chlorosis and bacterial specks expanded to the right side of it. The control plants where 10 mM of sucrose was injected without bacterial cells did not show any evident changes in the treated leaves (Fig. 7A). Treatment of 2.5 mM of 4-hydroxybenzoic acid and vanillic acid significantly abated disease symptoms (Fig. 7C and D).
We further investigated the effect of phenolic acids on bacterial population in tomato leaves. 25 ╬╝g of phenolic acids contained 17.8 ╬╝g of 4-hydroxybenzoic acid and 7.2 ╬╝g of vanillic acid of which quantity was 4 times higher in 100 ╬╝g of phenolic acids. As the amount of phenolic acids increased, cfu value remarkably decreased (Supplementary Fig. 2). The results suggested that phenolic acids would inhibit colonization of Pst DC3000 in tomato plants, and subsequent impact on bacterial survival.
Phenolic acids pervasive in a wide range of plants were known to protect the plants against UV light, invasion of insects or microorganisms and also have beneficial impacts on human health through diverse bioactivities such as antioxidant, antimicrobial and antitumor. Phenolic acids are produced from shikimate pathway from L-phenylalanine or L-tyrosine as the common precursors (Rice-Evans et al., 1996). Deamination of the two amino acids occurs to give cinnamic acid or p-coumaric acid, and then subsequent hydroxylation and methylation produce benzoic acid and phenolic derivatives (Rice-Evans et al., 1996). The most common parent structures of phenolic acid are hydroxybenzoic acid and hydroxycinnamic acid serving as the building blocks to form other plant natural products like sugars, organic acids and polymers (Russell and Duthie, 2011). For example, pectin is cross-linked to other polysaccharides through ester linkage between diferulic acid, which is formed from dimerization of hydroxycinnamic acids (Fry, 1986).
4-Hydroxybenzoic acid is one of the most abundant phenolic acids found in plants. It is reported to have antioxidant activity and antibacterial activities against various gram-negative and gram-positive bacteria (Cho et al., 1998; Rice-Evans et al., 1996). Also, 4-hydroxybenzoic acid is accumulated in plants with SA in response to a mobile signal induced by HR-causing pathogen, but the physiological role of it is not clear (Smith-Becker et al., 1998). Vanillic acid is also prevalent phenolic acid in plants along with 4-hydroxybenzoic acid to be found as an intermediate to produce vanillin from ferulic acid. In addition to antioxidant property, vanillic acid was reported to regulate inflammation by suppressing the activation of transcription factor which is crucial in immune responses (Kim et al., 2011). Furthermore, it was reported that virulence factors of Serratia marcescens, an opportunistic human pathogen and QS-regulated biofilm production were modulated by vanillic acid (Sethupathy et al., 2017).
In our studies, 4-hydroxybenzoic acid and vanillic acid were identified from S. middendorffianum to have inhibitory effect on the expression of T3SS in Pst DC3000. Both compounds downregulated the expression of GFP gene under the control of hrpA gene promoter regulating the expression of structural protein HrpA responsible for the construction of Hrp pili in Pst DC3000. Treatment of 4-hydroxybenzoic acid and vanillic acid also affected transcriptional level of other genes in hrp/hrc gene cluster, where hrpL and hopP1 genes were significantly downregulated. The hopP1 gene possesses hrp box sequence at the promoter region, which is recognized by HrpL, alternative sigma factor to activate the transcription of the gene. The characteristics of HopP1 are determined by two domains. One is harpin-like domain at N-terminal region which is similar to the harpin domain of HrpW1 and the other is soluble lytic transglycosylase domain, the fact of which implies that HopP1 might interact with peptidoglycan of the bacteria (Kvitko et al., 2007; Oh et al., 2007). The function of HopP1 is to elicit hypersensitive response in tobacco plants when it was delivered to the apoplast of the leaves (Kvitko et al., 2007). The atypical property of HopP1 is that it was only found in Pst DC3000, whereas other harpins such as HrpZ1, HrpW1, and HopAK1 were widely found in P. syringae pathovars (Kvitko et al., 2007). It has been reported that the suppression of hrpL transcription resulted in the decrease in the transcription of hrpL-dependent genes (Fouts et al., 2002). For that reason, although 4-hydroxybenzoic acid and vanillic acid were initially screened to inhibit the promoter activity of hrpA gene, the decrease in the transcription of hrpL gene might lead to reduction of hrpA and hopP1 transcripts. In recent study on genome-wide approach, 73 genes were identified to belong to new additional member regulated by HrpL, which suggests that other HrpL-dependent genes may also be downregulated by two phenolic acids (Fouts et al., 2002).
It was previously reported that some phenolic compounds including ortho-coumaric acid, trans-2-phenylcyclopropane-1-carboxylic acid, trans-2-methoxycinnamic acid and trans-2-methylcinnamic acid, inhibited the promoter activity of harpin gene in X. oryzae pv. oryzae which is the causal agent of bacterial blight in rice (Fan et al., 2017). Further experiments revealed that they attenuated hypersensitive response caused by X. oryzae pv. oryzae on tobacco leaves and suppressed the expression of genes located in hrp cluster. A simple phenolic acid, p-coumaric acid manipulated the expression of a core regulator of T3SS, HrpX/Y twocomponent system in Dickeya dadantii (Li et al., 2009). The activity of the compound was predicted to come from the position of the hydroxyl group and its double bond. The compound of which double bond is reduced, did not inhibit the expression of hrpA gene (Li et al., 2009). In the case of 4-hydroxybenzoic acid, 200 ╬╝M of the compound facilitated the accumulation of AvrPto, of which transcription is regulated by HrpL and secretion is dependent on Hrp pili of Pst DC3000 (Anderson et al., 2014). However, in our result, 4-hydroxybenzoic acid at 2.5 mM decreased the transcription of hrpL and hrpA genes. At the concentration of 200 ╬╝M, the promoter activity of hrpA gene was not affected by 4-hydroxybenzoic acid (data not shown). These results suggested that phenolic acids from natural sources possess ability to modulate the virulence and pathogenicity of plant pathogens and some of them may have potential to control plant diseases as anti-virulence agents.
In summary, 4-hydroxybenzoic acid and vanillic acid were identified from the extract of S. middendorffianum which was selected from T3SS inhibitor screening system. The transcription of three genes involved in T3SS were downregulated by 4-hydroxybenzoic acid and vanillic acid and they attenuated disease symptoms in tomato leaves infected by Pst DC3000. Even further experimental evidence is required to explicate the roles of phenolic acids in inhibition of T3SS, our results demonstrate the possibility of phenolic acids as anti-virulence agents to control bacterial diseases.
Supplementary materials are available at The Plant Pathology Journal website (http://www.ppjonline.org/).
Acknowledgments
This work was supported by the National Research Foundation of Korea (NRF) grant funded by the Korea Government (NRF-2017R1D1A1B03027996). The authors appreciate the support of an internal grant of Korea University.
Fig.┬Ā1
Type III secretion system inhibitory activity of plant extracts. Fluorescence intensity of strain CA9-5 treated with shoot and root extracts of Sedum middendorffianum and S. kamtschaticum. Asterisks indicate statistically significant differences (*P < 0.05).
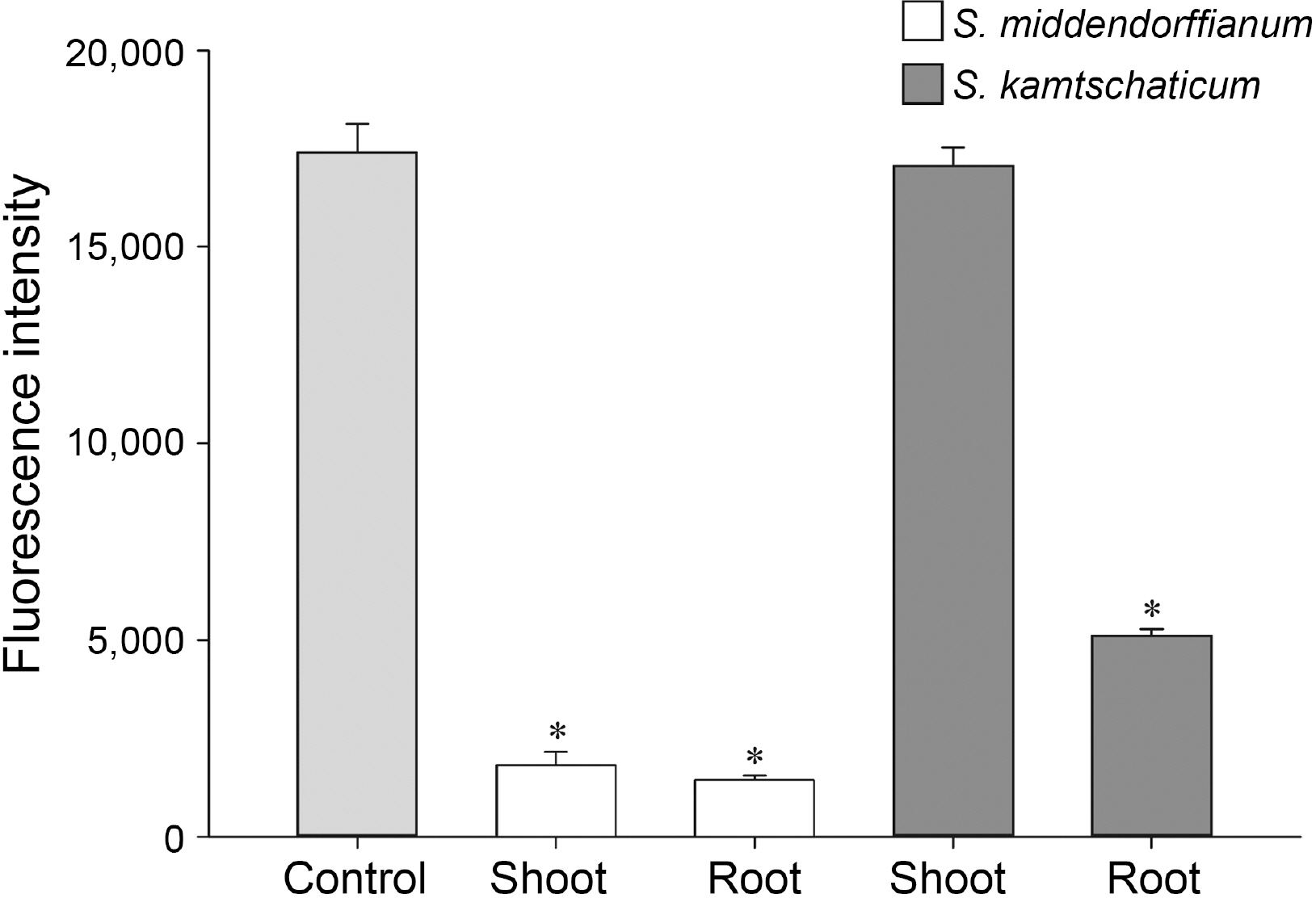
Fig.┬Ā2
Structures of 4-hydroxybenzoic acid (A) and vanillic acid (B) isolated from Sedum middendorffianum.
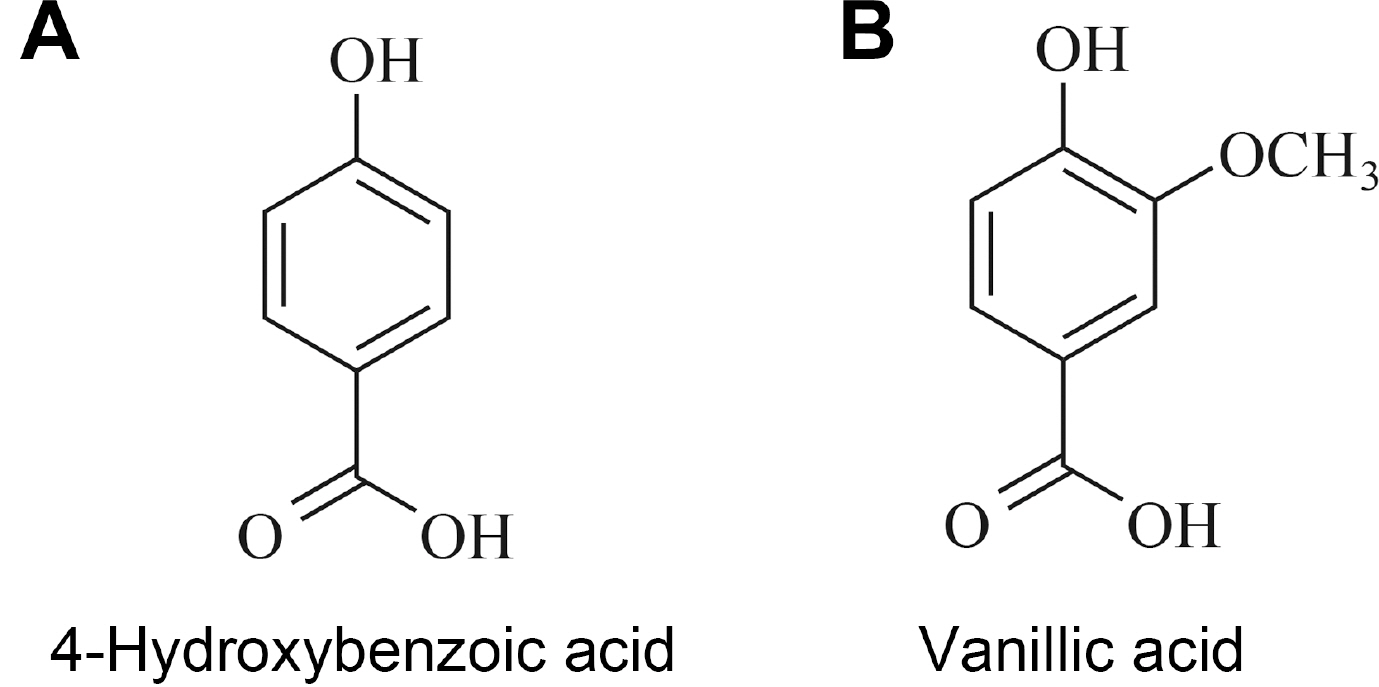
Fig.┬Ā3
Growth curve analysis of Pseudomonas syringae pv. tomato DC3000 treated with 4-hydroxybenzoic acid and vanillic acid. Optical density (OD600nm) of bacterial culture was measured every 2 h until 12 h after starting incubation and then every 4 h after peaking at the onset of the stationary phase. Each point indicates the average value of OD600nm of three replicates with error bar representing the standard deviation.
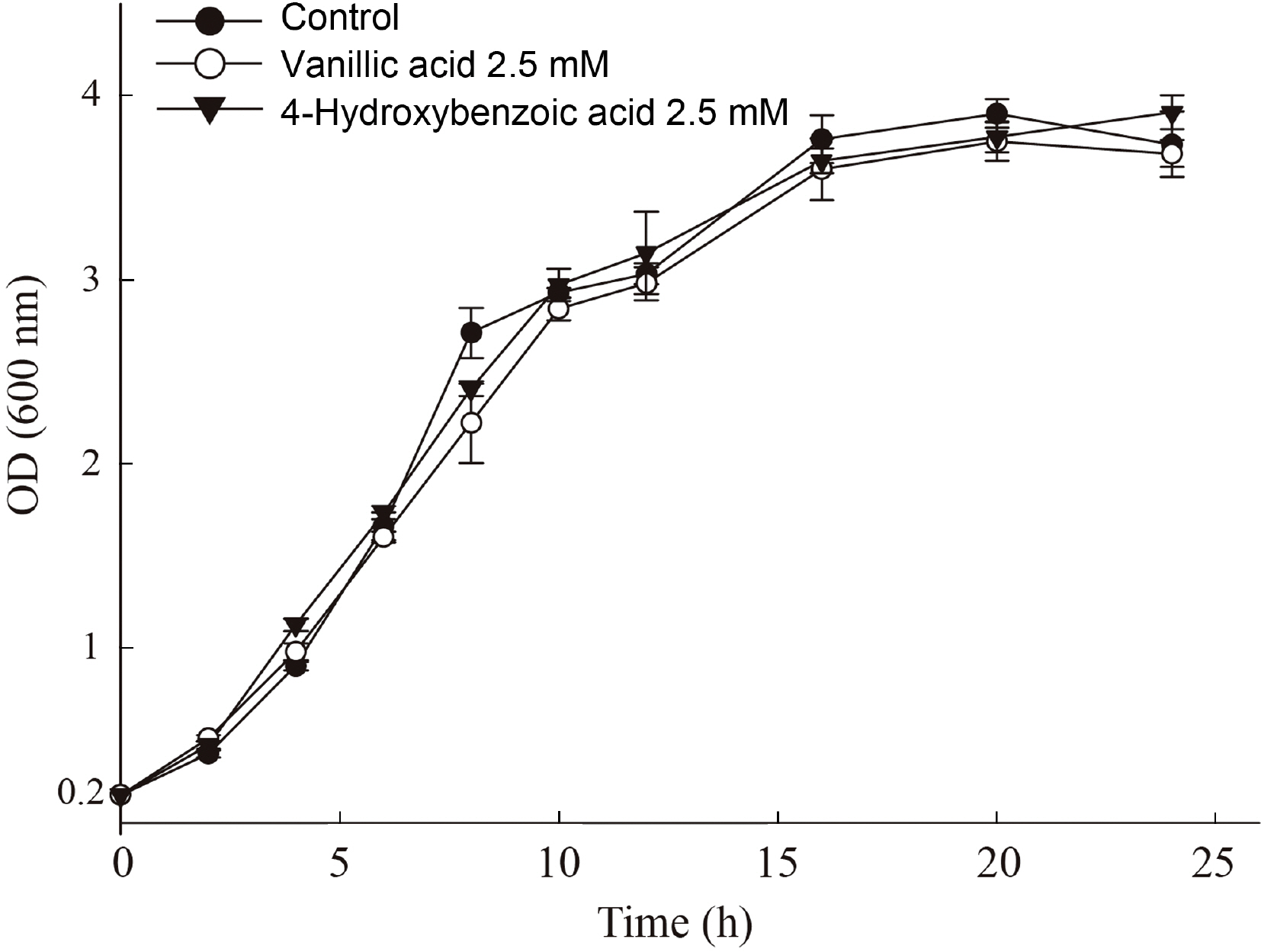
Fig.┬Ā4
Inhibitory effect of 4-hydroxybenzoic acid and vanillic acid on GFP-expressing Pseudomonas syringae pv. tomato DC3000. CA9-5 cells supplemented with two-fold serial dilutions of 4-hydroxybenzoic acid (A) and vanillic acid (B) were incubated in hrp--inducing medium. After 6 h, the fluorescence intensity of each treatment was measured. Different letters on the bars indicate significant difference (P < 0.05) between concentrations.
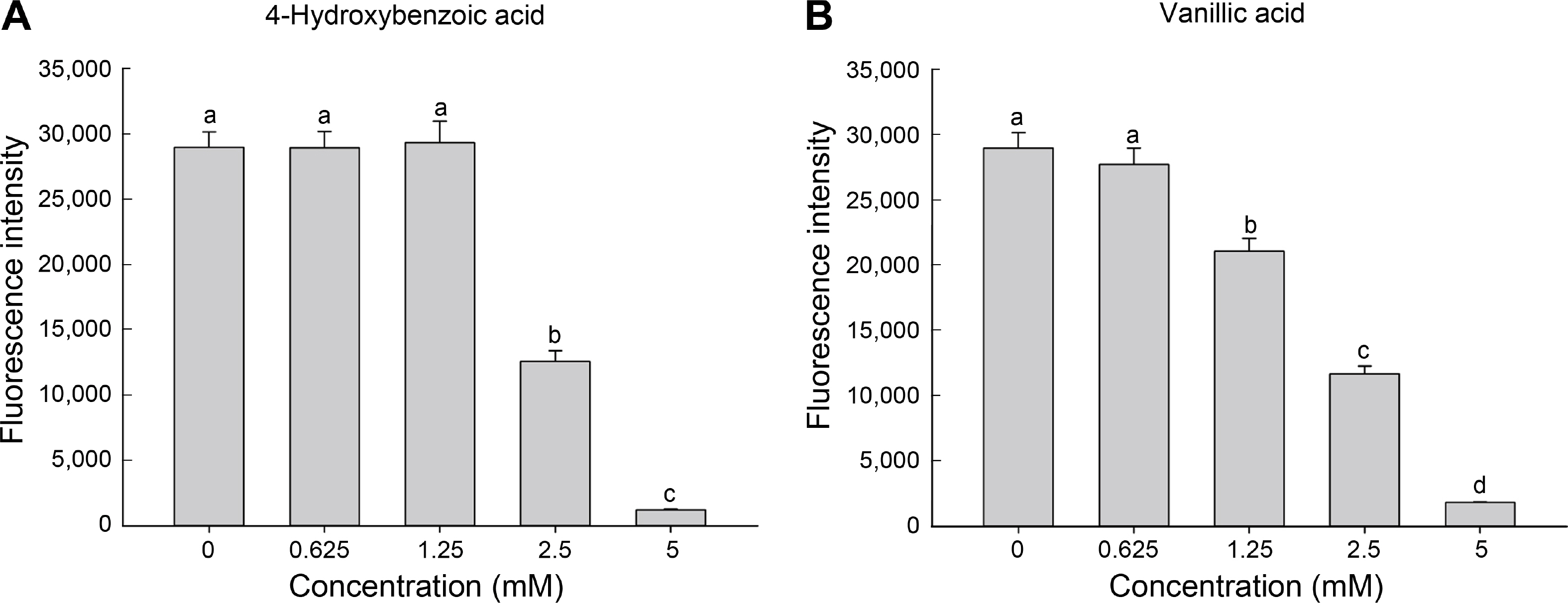
Fig.┬Ā5
Expression levels of type III secretion system (T3SS) genes. Transcript levels of four genes (gyrA, hopP1, hrpA and hrpL) of Pseudomonas syringae pv. tomato DC3000 treated with 2.5 mM of 4-hydroxybenzoic acid and vanillic acid were normalized by the expression of internal control gene, recA. The data is representative of three independent experiments with three replicates. Asterisks on the bar indicate significant difference (*P < 0.05).

Fig.┬Ā6
Auto-agglutination assay. Agglutinates of GFP-expressing Pseudomonas syringae pv. tomato DC3000 treated with 0.5% dimethyl sulfoxide (A), 4-hydroxybenzoic acid (B), and vanillic acid (C) were observed by fluorescence microscopy after 48 h incubation
in hrp-inducing medium. Scale bars = 10 ╬╝m.
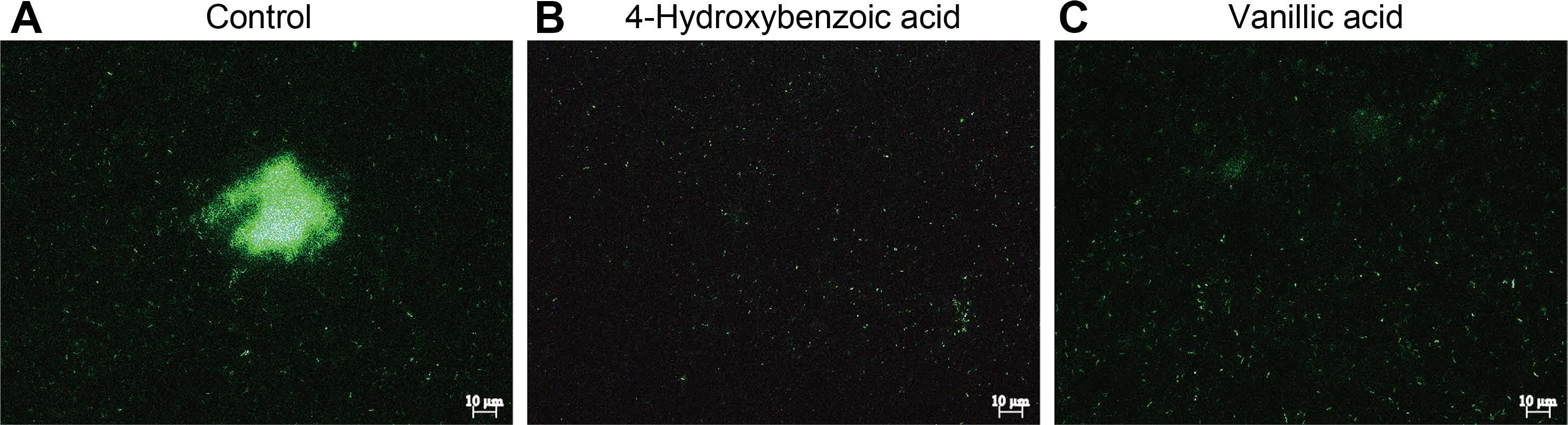
Fig.┬Ā7
Control efficacy of 4-hydroxybenzoic acid and vanillic acid on bacterial speck disease on tomato leaves. Ten mM of sucrose (A), Pst DC3000 suspension (5 ├Ś 103 cfu/ml) (B), and Pst DC3000 suspension with 2.5 mM of 4-hydroxybenzoic acid (C) and vanillic acid (D) were syringe infiltrated into the abaxial side of tomato leaves. Disease symptoms were observed 7 days after infection.

Table┬Ā1
Minimum inhibitory concentrations (MICs) of 4-hydroxybenzoic acid and vanillic acid against various plant pathogens
| Plant pathogenic microorganism | MICsa (mM) of phenolic acids | |
|---|---|---|
|
|
||
| 4-Hydroxybenzoic acid | Vanillic acid | |
| Pseudomonas syringae pv. tomato DC3000 | 10 | 10 |
| Erwinia carotovora subsp. carotovora ATCC39048 | 10 | 10 |
| Erwinia carotovora subsp. atroseptica BAA672 | 10 | 5 |
| Xanthomonas campestris pv. vesicatoria Ds1 | 10 | 10 |
| Fusarium oxysporum f. sp. lycopersici | >10b | 10 |
| Colletotrichum orbiculare | >10 | 10 |
REFERENCES
Alfano, J. R., Charkowski, A. O., Deng, W. L., Badel, J. L., Petnicki-Ocwieja, T., van Dijk, K. and Collmer, A. 2000. The Pseudomonas syringae Hrp pathogenicity island has a tripartite mosaic structure composed of a cluster of type III secretion genes bounded by exchangeable effector and conserved effector loci that contribute to parasitic fitness and pathogenicity in plants. Proc. Natl. Acad. Sci. U. S. A. 97:4856-4861.



Anderson, J. C., Wan, Y., Kim, Y.-M., Pasa-Tolic, L., Metz, T. O. and Peck, S. C. 2014. Decreased abundance of type III secretion system-inducing signals in Arabidopsis mkp1 enhances resistance against Pseudomonas syringae. Proc. Natl. Acad. Sci. U. S. A. 111:6846-6851.



Boureau, T., Routtu, J., Roine, E., Taira, S. and Romantschuk, M. 2002. Localization of hrpA-induced Pseudomonas syringae pv. tomato DC3000 in infected tomato leaves. Mol. Plant Pathol. 3:451-460.


Burr, T. J., Norelli, J. L., Katz, B., Wilcox, W. F. and Hoying, S. A. 1988. Streptomycin resistance of Pseudomonas syringae pv. papulans in apple orchards and its association with a conjugative plasmid. Phytopathology. 78:410-413.

Chiou, C.-S. and Jones, A. L. 1995. Molecular analysis of highlevel streptomycin resistance in Erwinia amylovora. Phytopathology. 85:324-328.

Cho, J.-Y., Moon, J.-H., Seong, K.-Y. and Park, K.-H. 1998. Antimicrobial activity of 4-hydroxybenzoic acid and trans 4-hydroxycinnamic acid isolated and identified from rice hull. Biosci. Biotechnol. Biochem. 62:2273-2276.


Dickey, S. W., Cheung, G. Y. C. and Otto, M. 2017. Different drugs for bad bugs: antivirulence strategies in the age of antibiotic resistance. Nat. Rev. Drug Discov. 16:457-471.


Fan, S., Tian, F., Li, J., Hutchins, W., Chen, H., Yang, F., Yuan, X., Cui, Z., Yang, C.-H. and He, C. 2017. Identification of phenolic compounds that suppress the virulence of Xanthomonas oryzae on rice via the type III secretion system. Mol. Plant Pathol. 18:555-568.


Fouts, D. E., Abramovitch, R. B., Alfano, J. R., Baldo, A. M., Buell, C. R., Cartinhour, S., Chatterjee, A. K., Ascenzo, M., Gwinn, M. L., Lazarowitz, S. G., Lin, N. C., Martin, G. B., Rehm, A. H., Schneider, D. J., van Dijk, K., Tang, X. and Collmer, A. 2002. Genomewide identification of Pseudomonas syringae pv. tomato DC3000 promoters controlled by the HrpL alternative sigma factor. Proc. Natl. Acad. Sci. U. S. A. 99:2275-2280.



Fry, S. C. 1986. Cross-linking of matrix polymers in the growing cell walls of angiosperms. Annu. Rev. Plant Physiol. 37:165-186.

Huynh, T. V., Dahlbeck, D. and Staskawicz, B. J. 1989. Bacterial blight of soybean: regulation of a pathogen gene determining host cultivar specificity. Science. 245:1374-1377.


Jagadeeswaran, G., Raina, S., Acharya, B. R., Maqbool, S. B., Mosher, S. L., Appel, H. M., Schultz, J. C., Klessig, D. F. and Raina, R. 2007. Arabidopsis GH3-LIKE DEFENSE GENE 1 is required for accumulation of salicylic acid, activation of defense responses and resistance to Pseudomonas syringae. Plant J. 51:234-246.


Kang, J. E., Han, J. W., Jeon, B. J. and Kim, B. S. 2016. Efficacies of quorum sensing inhibitors, piericidin A and glucopiericidin A, produced by Streptomyces xanthocidicus KPP01532 for the control of potato soft rot caused by Erwinia carotovora subsp. atroseptica. Microbiol. Res. 184:32-41.


Kang, J. E., Jeon, B. J., Park, M. Y., Yang, H. J., Kwon, J., Lee, D. H. and Kim, B. S. 2020. Inhibition of the type III secretion system of Pseudomonas syringae pv. tomato DC3000 by resveratrol oligomers identified in Vitis vinifera L. Pest Manag. Sci. 76:2294-2303.


Kim, J. D., Jeon, B. J., Han, J. W., Park, M. Y., Kang, S. A. and Kim, B. S. 2016. Evaluation of the endophytic nature of Bacillus amyloliquefaciens strain GYL4 and its efficacy in the control of anthracnose. Pest Manag. Sci. 72:1529-1536.


Kim, M.-C., Kim, S.-J., Kim, D.-S., Jeon, Y.-D., Park, S. J., Lee, H. S., Um, J.-Y. and Hong, S.-H. 2011. Vanillic acid inhibits inflammatory mediators by suppressing NF-╬║B in lipopolysaccharide-stimulated mouse peritoneal macrophages. Immunopharmacol. Immunotoxicol. 33:525-532.


King, E. O., Ward, M. K. and Raney, D. E. 1954. Two simple media for the demonstration of pyocyanin and fluorescin. J. Lab. Clin. Med. 44:301-307.

Kvitko, B. H., Ramos, A. R., Morello, J. E., Oh, H.-S. and Collmer, A. 2007. Identification of harpins in Pseudomonas syringae pv. tomato DC3000, which are functionally similar to HrpK1 in promoting translocation of type III secretion system effectors. J. Bacteriol. 189:8059-8072.



Leben, C. and Keitt, G. W. 1954. Antibiotics and plant disease, effects of antibiotics in control of plant diseases. J. Agric. Food Chem. 2:234-239.

Li, Y., Peng, Q., Selimi, D., Wang, Q., Charkowski, A. O., Chen, X. and Yang, C.-H. 2009. The plant phenolic compound pcoumaric acid represses gene expression in the Dickeya dadantii type III secretion system. Appl. Environ. Microbiol. 75:1223-1228.


Morgan, J. M., Duncan, M. C., Johnson, K. S., Diepold, A., Lam, H., Dupzyk, A. J., Martin, L. R., Wong, W. R., Armitage, J. P., Linington, R. G. and Auerbuch, V. 2017. Piericidin A1 blocks Yersinia Ysc type III secretion system needle assembly. mSphere. 2:e00030-17.




Newman, D. J. and Cragg, G. M. 2020. Natural products as sources of new drugs over the nearly four decades from 01/1981 to 09/2019. J. Nat. Prod. 83:770-803.


Oh, H.-S., Kvitko, B. H., Morello, J. E. and Collmer, A. 2007. Pseudomonas syringae lytic transglycosylases coregulated with the type III secretion system contribute to the translocation of effector proteins into plant cells. J. Bacteriol. 189:8277-8289.



Raskin, I. 1992. Role of salicylic acid in plants. Annu. Rev. Plant Physiol. Plant Mol. Biol. 43:439-463.

Rice-Evans, C. A., Miller, N. J. and Paganga, G. 1996. Structureantioxidant activity relationships of flavonoids and phenolic acids. Free Radic. Biol. Med. 20:933-956.


Russell, W. and Duthie, G. 2011. Plant secondary metabolites and gut health: the case for phenolic acids. Proc. Nutr. Soc. 70:389-396.


Ryals, J. A., Neuenschwander, U. H., Willits, M. G., Molina, A., Steiner, H.-Y. and Hunt, M. D. 1996. Systemic acquired resistance. Plant Cell. 8:1809-1819.



Sethupathy, S., Ananthi, S., Selvaraj, A., Shanmuganathan, B., Vigneshwari, L., Balamurugan, K., Mahalingam, S. and Pandian, S. K. 2017. Vanillic acid from Actinidia deliciosa impedes virulence in Serratia marcescens by affecting Slayer, flagellin and fatty acid biosynthesis proteins. Sci. Rep. 7:16328




Smith-Becker, J., Marois, E., Huguet, E. J., Midland, S. L., Sims, J. J. and Keen, N. T. 1998. Accumulation of salicylic acid and 4-hydroxybenzoic acid in phloem fluids of cucumber during systemic acquired resistance is preceded by a transient increase in phenylalanine ammonia-lyase activity in petioles and stems. Plant Physiol. 116:231-238.



Spotts, R. A. and Cervantes, L. A. 1995. Copper, oxytetracycline, and streptomycin resistance of Pseudomonas syringae pv syringae strains from pear orchards in Oregon and Washington. Plant Dis. 79:1132-1135.

- TOOLS
-
METRICS

- Related articles
-
Role of
Metcalfa pruinosa as a Vector forPseudomonas syringae pv.actinidiae 2017 December;33(6)




 PDF Links
PDF Links PubReader
PubReader ePub Link
ePub Link Full text via DOI
Full text via DOI Full text via PMC
Full text via PMC Download Citation
Download Citation Supplement
Supplement Print
Print