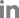|
 |
| Plant Pathol J > Volume 39(1); 2023 > Article |
|
Abstract
Phytoplasmas were discovered in diseased Elaeocarpus sylvestris trees growing on Jeju Island that showed symptoms of yellowing and darkening in the leaves. Leaf samples from 14 symptomatic plants in Jeju-si and Seogwipo-si were collected and phytoplasma 16S rRNA was successfully amplified by nested polymerase chain reaction using universal primers. The sequence analysis detected two phytoplasmas, which showed 99.5% identity to ‘Candidatus Phytoplasma asteris’ and ‘Ca. P. malaysianum’ affiliated to 16SrI and 16SrXXXII groups, respectively. Through polymerase chain reaction-restriction fragment length polymorphism (RFLP) analyses using the AfaI (RsaI) restriction enzyme, the presence of two phytoplasmas strains as well as cases of mixed infection of these strains was detected. In a virtual RFLP analysis with 17 restriction enzymes, the 16S rRNA sequence of the ‘Ca. P. asteris’ strain was found to match the pattern of the 16SrI-B subgroup. In addition, the phytoplasmas in the mixed-infection cases could be distinguished using specific primer sets. In conclusion, this study confirmed mixed infection of two phytoplasmas in one E. sylvestris plant, and also the presence of two phytoplasmas (of the 16SrI and 16SrXXXII groups) in Jeju Island (Republic of Korea).
Phytoplasmas are pleomorphic bacteria 80-800 nm in size that possess a very small genome of about 680-1,600 kb (Pagliari and Musetti, 2019). They are plant pathogens transmitted primarily by phloem-feeding leaf hoppers and plant hoppers (Weintraub and Beanland, 2006; Weintraub and Jones, 2009) and are known to affect over 1,000 species worldwide. Infected plants show various symptoms, including virescence, phyllody, shortened internodes, dwarfed leaves, witches’ broom, flower malformation, yellowing, and decline (Bertaccini et al., 2014; Kumari et al., 2019; Namba, 2019). Phytoplasmas can be detected via amplification of the 16S rRNA gene, and classified into ribosomal groups by restriction fragment length polymorphism (RFLP) and sequencing of 16S rDNA (Lee et al., 1998; Lim and Sears, 1989).
Elaeocarpus sylvestris var. ellipticus (Thunb.) H. Hara is a common tree that constitutes 5% of all trees in the streets of South Korea’s Jeju Island. Since 2013, E. sylvestris trees have been observed to die after developing symptoms of E. sylvestris decline, which has been associated with the presence of ‘Candidatus Phytoplasma malaysianum’ (16SrXXXII) in South Korea (Lee et al., 2022). This phytoplasma has been reported also as associated with the death of Elaeocarpus trees in Japan since the 1980s (Iwabuchi et al., 2018), and another phytoplasma has been identified as associated with Elaeocarpus zollingeri yellows (Kawabe et al., 2000; Satoh et al., 2014). Several studies have reported ‘Ca. P. malaysianum’ and ‘Ca. P. asteris’ presence in E. zollingeri trees in Japan (Iwabuchi et al., 2018; Kawabe et al., 2011; Satoh et al., 2014). Considering that the symptoms were similar to those of E. zollingeri infected with phytoplasmas, it was presumed that phytoplasmas are present in E. sylvestris. Therefore, the purpose of this study was to screen for the presence of phytoplasmas in E. sylvestris declining trees of Jeju Island in the Republic of Korea.
Between 2021 and 2022, E. sylvestris were monitored for decline symptoms. Samples were collected from seven E. sylvestris in Jeju-si and another seven E. sylvestris in Seogwipo-si that showed symptoms of yellowing and darkening leaves. Asymptomatic trees were collected from two E. sylvestris in both regions, respectively. Genomic DNA was extracted from 0.2 g of vein tissue in fresh leaves from each sample using a DNeasy Plant Mini Kit (Qiagen, Hilden, Germany).
Phytoplasma detection was performed by polymerase chain reaction (PCR) using universal primer set P1 and P7. Nested PCR was conducted using the primers R16F2n and R16R2, and the amplification products were diluted in distilled water (v/v, 1:30) (Deng and Hiruki, 1991; Gundersen and Lee, 1996; Lee et al., 1993; Schneider et al., 1995). To distinguish phytoplasmas in cases of mixed infection, new specific primers were designed to detect phytoplasma strains in the 16SrXXXII group: R16(XXXII)F1 (5′-CGTGGGTAGCAAACAGGATT-3′)/R16(XXXII)R1 (5′-GCCACTGCTTTCGGGTATTA-3′). To detect phytoplasmas in the 16SrI group, R16(I)F1 and R16(I)R1 primers were used (Lee et al., 1994). To identify the ribosomal protein (rp) operon and tuf genes, the universal primers rpF1/rpR1 (Lee et al., 1998) and fTuf1/rTuf1 were used, respectively (Schneider and Gibb, 1997). The PCR reaction mixture (total volume; 25 μl) consisted of 50-100 ng of genomic DNA, 15 μl of Emerald-Amp GT PCR Master Mix [2× Premix] (TaKaRa, Shiga, Japan), and 10 pmol of each forward and reverse primer, with the remaining volume being distilled water. The conditions for PCR amplification were as follows: one cycle of denaturation at 94°C for 7 min; 35 cycles (28 cycles for the nested PCR) of denaturation at 94°C for 1 min, annealing at primer-specific temperatures (P1/P7: 55°C, R16F2n/R2: 58°C, R16(XXXII)F1/R16(XXXII)R1: 59°C, R16(I)F1/R16(I)R1: 53°C, rpF1/rpR1: 53°C, and fTuf1/rTuf1: 56°C) for 2 min, and extension at 72°C for 3 min; and a final extension step of 72°C for 10 min.
The secA gene was detected using the phytoplasma universal primers SecAfor1 and SecArev3. The PCR cycling conditions were modified as follows: a denaturation step for 2 min at 94°C; 30 total cycles at 94°C for 30 s, 55°C for 60 s, and 72°C for 90 s; and a final extension step at 72°C for 15 min (Hodgetts et al., 2008). The rp operon, tuf, and secA gene were amplified and sequenced using primers for rpF1/rpR1 (1.24 kb), fTuf1/rTuf1 (1.08 kb), and SecAfor1/SecArev3 (0.84 kb). To visualize amplification products, each PCR product was mixed with Dyne Loading STAR (Dyne, Seoul, Korea), and then separated on 1.2% agarose gels in TAE (Tris-Acetate-EDTA) buffer and visualized using a UV transilluminator. RFLP analysis was conducted with the restriction enzyme AfaI (RsaI) (TaKaRa, Otsu, Japan). The reaction was carried out using 450-500 ng of each PCR product, 10 U enzyme, and restriction digestion buffer (TaKaRa, Otsu, Japan). The mixture was incubated in a water bath at 37°C for 2 h. The PCR products and RFLP mixture were combined with Dyne Loading STAR, separated on 1.5% agarose gels in TAE buffer, and visualized using a UV transilluminator. Previously reported DNA of E. sylvestris decline phytoplasma strain ESDP-JJ1 (GenBank accession no. OL689203) was used as a positive control for RFLP analysis (Lee et al., 2022).
The amplified PCR product was purified using a PCR Purification Kit (Bioneer, Seoul, Korea). The sequencing analysis of 16S rRNA, 16-23S internal transcribed spacer region, 23S rDNA partial, rp, tuf, and secA amplicons was performed by Macrogen (Daejeon, Korea). The sequences were analyzed using MEGA 11.0 (Saitou and Nei, 1987), after which phytoplasma sequences were compared with those retrieved from GenBank, and phylogenetic analysis was carried out using the neighbor-joining system in MEGA 11.0 with default values and 10,000 bootstrap replicates (Tamura et al., 2021). The resulting 16S rRNA gene sequences were subjected to virtual RFLP analysis via the iPhyClassifier software (Zhao et al., 2009).
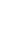
Since 2013, phytoplasma infection has been reported in E. sylvestris trees, a primary tree species in the streets of Jeju Island, with most trees dying due to ‘Ca. P. malaysianum’ presence (Lee et al., 2022). This phytoplasma has been associated with Malaysian periwinkle virescence, Malaysian oil palm, Malayan yellow dwarf, Camptotheca acuminate witches’ broom, Trema tomentosa witches’ broom (TtWB), and Elaeocarpus zollingeri yellows (Iwabuchi et al., 2018; Nejat et al., 2009, 2013; Yu et al., 2021). The main symptoms of decline in E. sylvestris trees are yellowing and darkening of the leaves; eventually, the leaves of infected trees drop and the leafless branches die (Fig. 1).
PCR amplification with P1/P7 primers detected the expected 1.83 kb product in all 14 samples. However, no bands appeared in samples collected from two asymptomatic trees. By performing the nested PCR reaction with these PCR products using the R16F2n/R2 primer set, 1.24 kb products were amplified. RFLP using the AfaI (RsaI) enzyme on 16S rRNA nested PCR products indicated three band patterns (Fig. 2). The pattern of group A was noticed in samples JJ011, JJ013, JJ014, JJ017, JS011, JS014, JS015, and JS016; while that of group B was detected in JJ012, JJ015, JS012, and JS013; and that of group C was found in JJ016 and JS017. The rp operon, tuf, and secA of group B samples were amplified to the expected band size and results showed 100% identity to each other and to phytoplasma strains belonging to ‘Ca. P. asteris’.
Based on the PCR-RFLP results, as mentioned above, group A samples showed the same band patterns as ‘Ca. P. malaysianum’ (Lee et al., 2022), group B samples appeared to be identical to ‘Ca. P. asteris’ (Lee et al., 1998). In addition, group C samples have both two phytoplasma fragment bands, therefore mixed infection of the two phytoplasmas was also detected.
Not including samples JJ016 and JS017, which showed evidence of a mixed infection, 12 samples exhibited a virtual RFLP pattern from BstUI, HhaI, BfaI, and key 16S rRNA restriction enzymes derived from the query 16S rDNA F2nR2 fragment that was identical to the 16SrI-B subgroup and 16SrXXXII group reference pattern (Figs. 3 and 4). When analyzing the four group B samples (JJ012, JJ015, JS012, and JS013) using iPhyClassifier (Zhao et al., 2009), the query 16S rDNA sequence showed 99.9% identity with that of ‘Ca. P. asteris’. In the 16S ribosomal group/subgroup classification, the pattern of 16S rDNA of E. sylvestris phytoplasmas was identical to that of 16Sr group I-B. In summary, the newly identified phytoplasma in E. sylvestris was determined to be a member of the 16SrI-B subgroup.
On the other hand, ‘Ca. P. malaysianum’ (16SrXXXII) phytoplasma presence in E. sylvestris showed a different pattern from that of 16SrXXXII-A, -B, -C, and -D subgroups in BstUI enzyme digestion, and could be distinguished from the 16SrXXXII-A subgroup in HhaI enzyme. ‘Ca. P. malaysianum’ was distinguished from the 16SrXXXII-B subgroup by Sau3AI enzyme, 16SrXXXII-C subgroup by BfaI enzyme, and 16SrXXXII-D subgroup by BstUI enzyme digestion (Nejat et al., 2009, 2013; Yu et al., 2021). Thus, this phytoplasma could be enclosed in another subgroup within the 16SrXXXII group tentatively named 16SrXXXII-E.
The group-specific PCR method applied here can be used to distinguish the presence of phytoplasmas in a mixed infection. Staniulis et al. (2000) and Alsaheli et al. (2020) accomplished this by using group-specific primers in plants coinfected with two phytoplasmas. Results of PCR using the group-specific primer sets R16(XXXII)F1/R16(XXXII)R1 (0.66 kb products) and R16(I)F1/R16(I)R1 (1.1 kb products) confirmed that the eight group A samples were infected with phytoplasma strains belonging to the ‘Ca. P. malaysianum’ (16SrXXXIIgroup); the four group B samples were infected with the ‘Ca. P. asteris’ (16SrI group); and the two group C samples were infected with both phytoplasmas (Table 1). Phylogenetic analysis of the phytopla-sma-related taxa based on 16S rDNA, rp operon, tuf, and secA sequences revealed that these regions share more than 99.5% similarity with those of ‘Ca. P. asteris’ group such as mulberry dwarf disease, sumac witches’ broom disease in South Korea caused by 16SrI-B group phytoplasma. Also, the phylogenetic classification of rp operon, tuf, and secA sequences is closely related 16SrI group phytoplasma (Figs. 5 and 6). The nucleotide sequences for the four genes were deposited in GenBank with the accession numbers: ON420422 and ON420423 (16S rDNA), ON420424 and ON420425 (secA gene), ON420426 and ON420427 (rp operon), and ON420428 and ON420429 (tuf gene).
In terms of decline symptoms, there was no difference between trees infected with 16SrI or 16SrXXXII groups, or in cases of a mixed infection. Several mixed-infection cases involving different phytoplasmas have been reported in clover (Staniulis et al., 2000), lily (Bertaccini et al., 2000), jujube tree (Lee et al., 2009), carrot (Mehle et al., 2018), and fig tree (Alsaheli et al., 2020). Currently, many E. sylvestris trees are dying due to phytoplasma infection in Jeju Island. In order to reduce the incidence of this disease, studies that examine insect vectors and the effect of trunk injection of antibiotics are required in the near future.
Acknowledgments
This paper was supported by a project, “Investigation of Diagnosis and Characteristics of Yellow Disease on Elaeocarpus sylvestris trees (Project No. FE0100-2018-10-2022)” from the National Institute of Forest Science, Republic of Korea.
Fig.¬Ý1
Various decline symptoms in Elaeocarpus sylvestris infected with phytoplasmas. (A) ‘Candidatus Phytoplasma malaysianum’ (16SrXXXII). (B) ‘Candidatus Phytoplasma asteris’ (16SrI). (C) Two phytoplasma mixed infection in a single E. sylvestris. (D) Asymptomatic tree (left), decline symptoms tree (right).
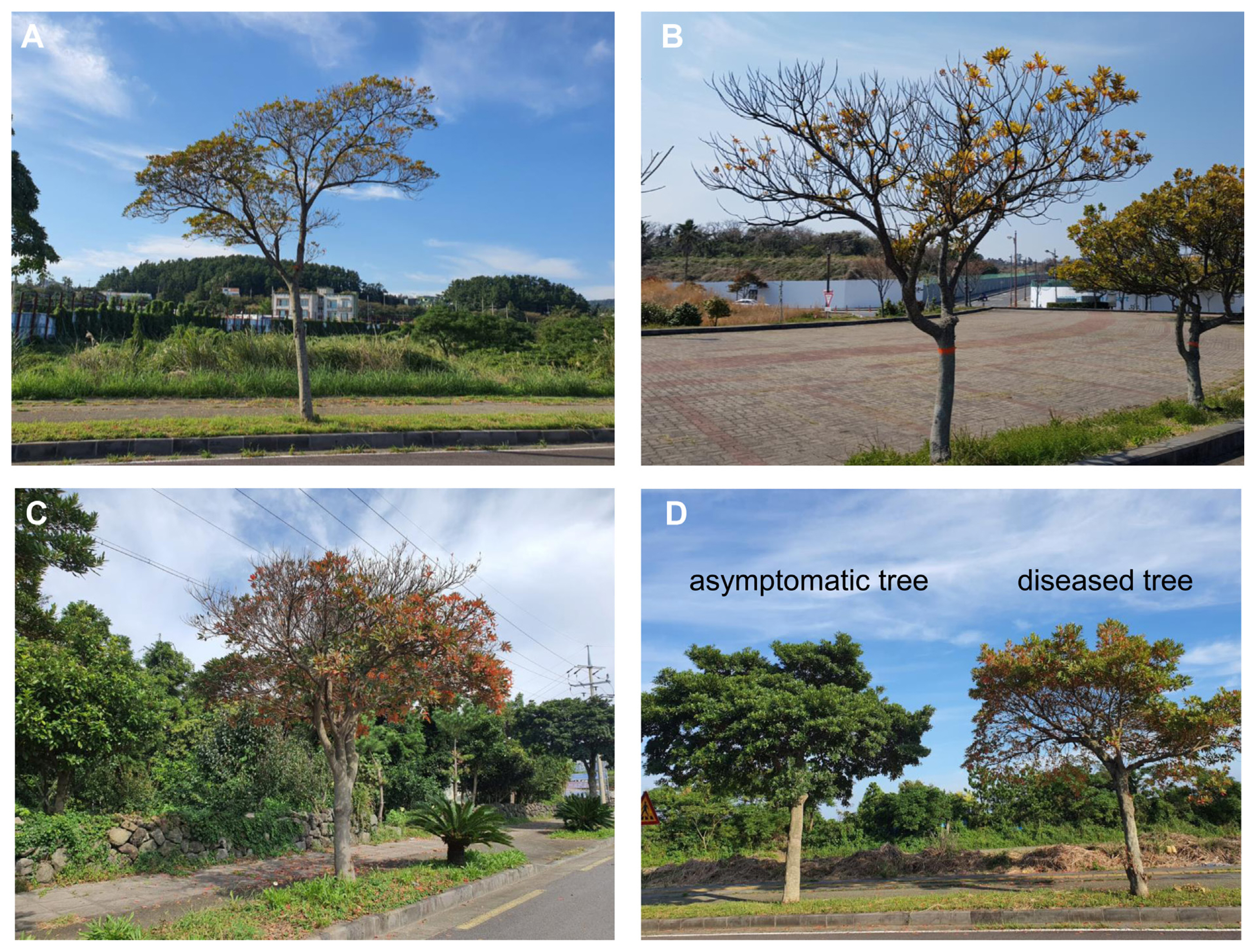

Fig.¬Ý2
Restriction fragment length polymorphism profiles of 16S rRNA gene amplified by nested polymerase chain reaction using primer pair R16F2n/R16R2 with digested AfaI (RsaI). Lane M, molecular weight marker (100 bp); lanes 1-7, JJ011-JJ017; lanes 8-14, JS011-JS017; lane 15, Elaeocarpus sylvestris, decline phytoplasma ESDP-JJ1 (GenBank accession no. OL689203).
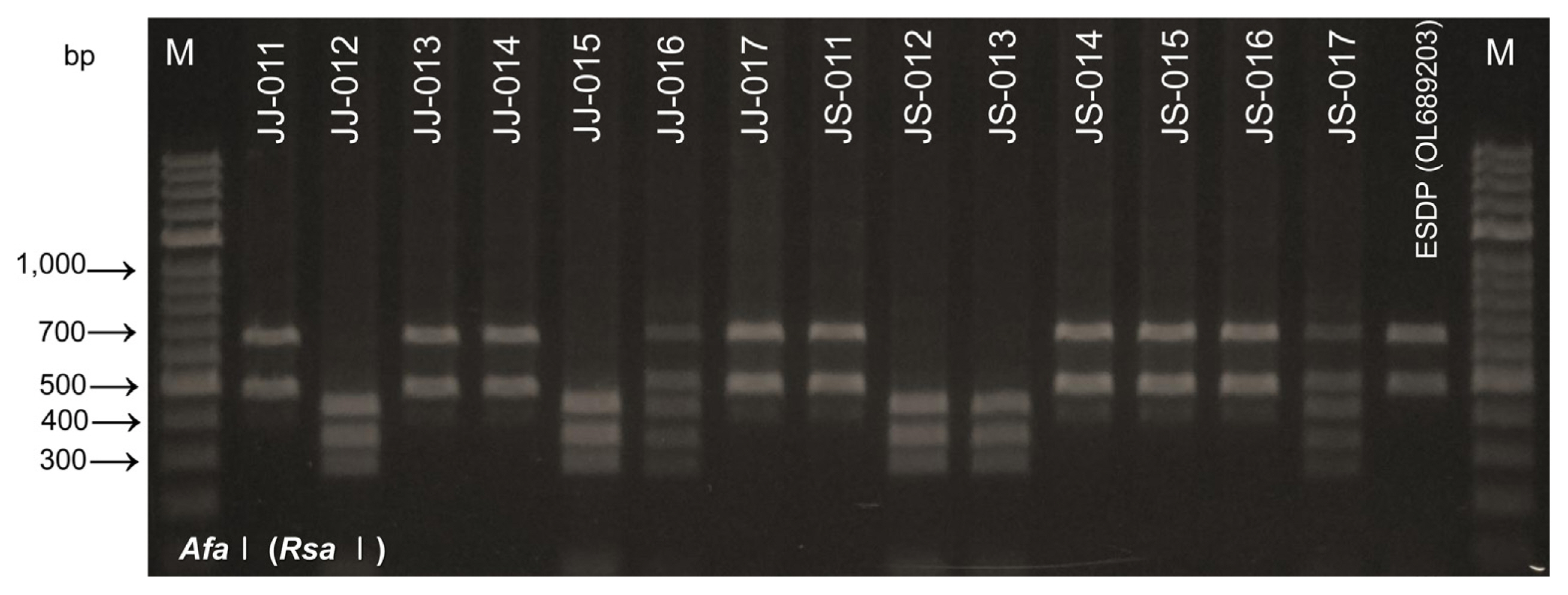
Fig.¬Ý3
Virtual restriction fragment length polymorphism (RFLP) profiles of phytoplasma associated with diseased tree in-silico digestions. Polymerase chain reaction products of JJ011, JJ012, JJ013, JJ014, JJ015, JJ017, and JS011, JS012, JS013, JS014, JS015, and JS016 were digested with restriction enzymes, BstUI (A), HhaI (B), BfaI (C), and Sau3AI (D) to compare the RFLP for each other. ESDP (16SrXXXII) GenBank accession no.: OL689203; Malaysian periwinkle virescence (MaPV; 16SrXXXII-A) GenBank accession no.: EU371934; Malayan yellow dwarf (MYD; 16SrXXXII-B) GenBank accession no.: EU498727; Malayan oil palm (MOP; 16SrXXXII-C) GenBank accession no.: EU498728; Trema tomentosa witches’ broom (TtWB; 16SrXXXII-D) GenBank accession no.: MW138004; MW: ΦX174DNA digested with HaeIII; fragment sizes 1,353, 1,078, 872, 603, 310, 281, 271, 234, 194, 118, and 72 base pairs, respectively.
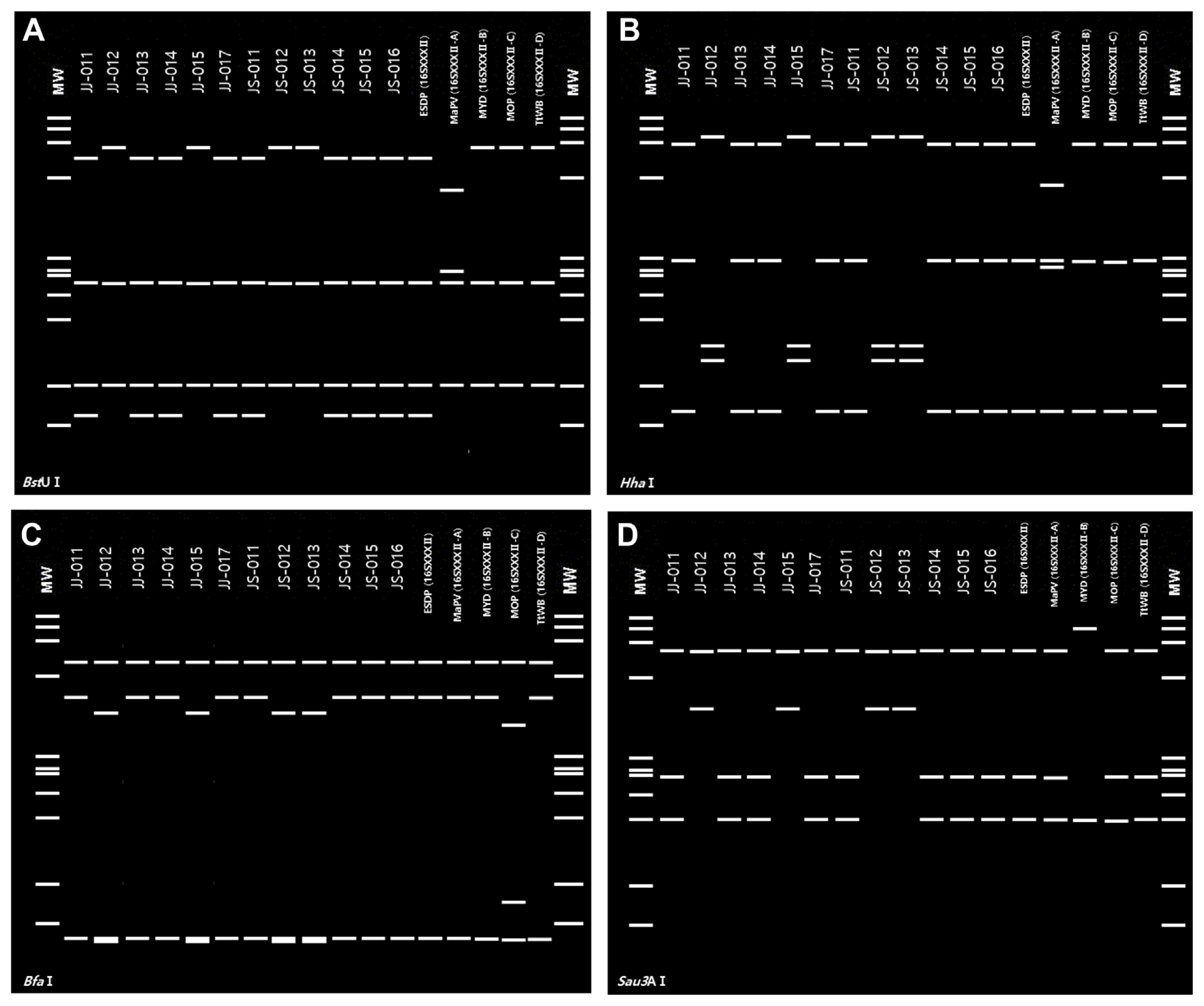

Fig.¬Ý4
Virtual restriction fragment length polymorphism (RFLP) profiles of phytoplasmas presence in Elaeocarpus sylvestris. ‘Candidatus P. malaysianum’ (GenBank accession no. OL689203) (A) and ‘Ca. P. asteris’ (GenBank accession no. ON420422) (B) were used with restriction 17 enzymes were used to compare the restriction fragment length polymorphism pattern results. MW, ΦX174DNA digested with HaeIII; fragment sizes 1,353, 1,078, 872, 603, 310, 281, 271, 234, 194, 118, and 72 base pairs, respectively.

Fig.¬Ý5
Phylogenetic relationship of two phytoplasmas in diseased tree from nucleotide sequences of 16S rRNA gene (combination of two phytoplasmas in Elaeocarpus sylvestris trees, ‘Candidatus P. malaysianum’ [GenBank accession no. OL689203-OL689207] and ‘Ca. P. asteris’ [GenBank accession no. ON420422-ON720423]) performed with MEGA 11.0 software using the neighbor-joining method with the default values and 1,000 replications for bootstrap analysis. Acholeplasma palmae was used as the out-group.
Fig.¬Ý6
Phylogenetic relationship of phytoplasmas from nucleotide sequences of secA gene (A) (GenBank accession no. ON420424-ON420425), rp operon (B) (GenBank accession no. ON420426-ON420427), tuf gene (C) (GenBank accession no. ON420428-ON420429), performed with MEGA 11.0 software using the neighbor-joining method with the default values and 1,000 replications for bootstrap analysis.

Table¬Ý1
Detection of phytoplasma 16S rRNA groups in Elaeocarpus sylvestris using group-specific primer sets and PCR-RFLP analyses
References
Alsaheli, Z., Contaldo, N., Mehle, N., Dermastia, M., Elbeaino, T. and Bertaccini, A. 2020. First detection of ‘Candidatus Phytoplasma asteris’ and ‘Candidatus Phytoplasma solani’-related strains in fig trees. J. Phytopathol 168:63-71.


Bertaccini, A., Botti, S., Martini, M. and Kaminska, M. 2000. Molecular evidence for mixed phytoplasma infection in lily plants. Acta Hortic 568:35-41.

Bertaccini, A., Duduk, B., Paltrinieri, S. and Contaldo, N. 2014. Phytoplasmas and phytoplasma diseases: a severe threat to agriculture. Am. J. Plant Sci 5:1763-1788.


Deng, S. and Hiruki, C. 1991. Amplification of 16S rRNA genes from culturable and nonculturable mollicutes. J. Microbiol. Methods 14:53-61.

Gundersen, D. E. and Lee, I.-M. 1996. Ultrasensitive detection of phytoplasmas by nested-PCR assays using two universal primer pairs. Phytopathol. Mediterr 35:144-151.
Hodgetts, J., Boonham, N., Mumford, R., Harrison, N. and Dickinson, M. 2008. Phytoplasma phylogenetics based on analysis of secA and 23S rRNA gene sequences for improved resolution of candidate species of ‘Candidatus Phytoplasma’. Int. J. Syst. Evol. Microbiol 58:1826-1837.


Iwabuchi, N., Endo, A., Kameyama, N., Satoh, M., Miyazaki, A., Koinuma, H., Kitazawa, Y., Maejima, K., Yamaji, Y., Oshima, K. and Namba, S. 2018. First report of ‘Candidatus Phytoplasma malaysianum’ associated with Elaeocarpus yellows of Elaeocarpus zollingeri
. J. Gen. Plant Pathol 84:160-164.


Kawabe, Y., Kusunoki, M., Miyashita, S. and Kikuchi, Y. 2000. Genetic diagnosis of phytoplasma diseases on trees. For. For. Prod. Res. Inst. Kenkyu Seika Senshu 2000:10-11 (in Japanese).
Kawabe, Y., Tsuda, J., Matsuura, K., Ogawa, S., Usami, Y. and Kusunoki, M. 2011. Analysis of oxytetracycline and detection of phytoplasma after the trunk injection of oxytetracycline formulation in Elaeocarpus sylvestris var. ellipticus infected with Elaeocarpus yellows. Tree For. Health 15:97-101.
Kumari, S., Nagendran, K., Rai, A. B., Singh, B., Rao, GP. and Bertaccini, A. 2019. Global status of phytoplasma diseases in vegetable crops. Front. Microbiol 10:1349.



Lee, G.-W., Han, T.-W., Lee, S. K. and Han, S.-S. 2022. ‘Candidatus Phytoplasma malaysianum’ (16SrXXXII) associated with Elaeocarpus sylvestris decline in South Korea. For. Sci. Technol 18:7-13.

Lee, I.-M., Gundersen-Rindal, D. E., Davis, R. E. and Bartoszyk, I. M. 1998. Revised classification scheme of phytoplasmas based on RFLP analyses of 16S rRNA and ribosomal protein gene sequences. Int. J. Syst. Bacteriol 48:1153-1169.

Lee, I.-M., Gundersen, D. E., Hammond, R. W. and Davis, R. E. 1994. Use of mycoplasmalike organism (MLO) group-specific oligonucleotide primers for nested-PCR assays to detect mixed-MLO infections in a single host plant. Phytopathology 84:559-566.

Lee, I.-M., Hammond, R. W., Davis, R. E. and Gundersen, D. E. 1993. Universal amplification and analysis of pathogen 16S rRNA for classification and identification of mycoplasmalike organisms. Mol. Plant Pathol 83:834-842.
Lee, S., Han, S. and Cha, B. 2009. Mixed infection of 16S rDNA I and V groups of phytoplasma in a single jujube tree. Plant Pathol. J 25:21-25.

Lim, P. O. and Sears, B .B. 1989. 16S rRNA sequence indicates that plant-pathogenic mycoplasmalike organisms are evolutionarily distinct from animal mycoplasmas. J. Bacteriol 171:5901-5906.




Mehle, N., Mermal, S., Vidmar, S., Marn, M. V., Dreo, T. and Dermastia, M. 2018. First report of carrot infection with phytoplasmas in Slovenia. In: Proceedings of the 5th European Bois Noir Workshop; 1-4; Ljubljana, Slovenia.
Namba, S. 2019. Molecular and biological properties of phytoplasmas. Proc. Jpn. Acad. Ser. B Phys. Biol. Sci 95:401-418.



Nejat, N., Sijam, K., Abdullah, S. N. A., Vadamalai, G. and Dickinson, M. 2009. Phytoplasmas associated with disease of coconut in Malaysia: phylogenetic groups and host plant species. Plant Pathol 58:1152-1160.

Nejat, N., Vadamalai, G., Davis, R. E., Harrison, N. A., Sijam, K., Dickinson, M., Abdullah, S. N. and Zhao, Y. 2013. ‘Candidatus Phytoplasma malaysianum’, a novel taxon associated with virescence and phyllody of Madagascar periwinkle (Catharanthus roseus). Int. J. Syst. Evol. Microbiol 63:540-548.


Saitou, N. and Nei, M. 1987. The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol. Biol. Evol 4:406-425.

Satoh, M., Takahashi, H., Chikamori, M., Tani, Y. and Adachi, N. 2014. Prevalence of Elaeocarpus yellows in Mt. Shiroyama in Tokushima City. Nat. Sci. Res. Univ. Tokushima 28:21-25.
Schneider, B. and Gibb, K. S. 1997. Sequence and RFLP analysis of the elongation factor Tu gene used in differentiation and classification of phytoplasmas. Microbiology 143:3381-3389.


Schneider, B., Seemuller, E., Smart, C. D. and Kirkpatrick, B. C. 1995. Phylogenetic classification of plant pathogenic mycoplasma-like organisms or phytoplasma. In: Molecular and diagnostic procedures in mycoplasmology, eds. by S. Razin and J. G. Tully, pp. 369-380. Academic Press, San Diego, CA, USA.
Staniulis, J. B., Davis, R. E., Jomantiene, R., Kalvelyte, A. and Dally, E. L. 2000. Single and mixed phytoplasma infections in phyllody- and dwarf-diseased clover plants in Lithuania. Plant Dis 84:1061-1066.


Tamura, K., Stecher, G. and Kumar, S. 2021. MEGA11: molecular evolutionary genetics analysis version 11. Mol. Biol. Evol 38:3022-3027.




Weintraub, P. G. and Beanland, L. 2006. Insect vectors of phytoplasmas. Annu. Rev. Entomol 51:91-111.


Weintraub, P. G. and Jones, P. 2009. Phytoplasmas: genomes, plant hosts and vectors. CABI International, Wallingford, UK. pp. 331.
- TOOLS
-
METRICS

-
- 0 Crossref
- 0 Scopus
- 1,445 View
- 102 Download
- ORCID iDs
-
Sang-Sub Han

https://orcid.org/0000-0002-1162-3110 - Related articles



 PDF Links
PDF Links PubReader
PubReader ePub Link
ePub Link Full text via DOI
Full text via DOI Full text via PMC
Full text via PMC Download Citation
Download Citation Print
Print