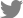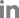|
 |
| Plant Pathol J > Volume 33(2); 2017 > Article |
Abstract
Tan spot, caused by the fungus Pyrenophora triticirepentis, is economically important foliar disease in Latvia, Lithuania, and Romania; however, race structure from Baltic States and Romania is not known. In this study, we performed genotypic and phenotypic race characterization of a large collection of P. triticirepentis isolates from these countries to determine race structure and utilize this information for better disease management and breeding wheat for tan spot resistance. We characterized 231 single spore isolates from Latvia (n = 15), Lithuania (n = 107), and Romania (n = 109) for Ptr ToxA and Ptr ToxB genes using two genes specific primers. A subset (139) of 231 isolates were further characterized for their race structure by inoculating them individually on tan spot wheat differentials set. Majority (83%) of the 231 isolates amplified Ptr ToxA gene suggesting prevalence of race 1 and 2. Further, phenotypic characterization of 139 isolates also showed wide prevalence of races 1 (68%), 2 (8%), 3 (11%), and 4 (5%) were also identified from Baltic States as well as Romania. Eighteen of the isolates (13%) did not seem to be of any of the eight known races as they lacked Ptr ToxA gene but they behaved like either race 1 or race 2, suggesting possibility of novel toxins in these isolates as their virulence tools.
Wheat is ranked first among small grains production in the Baltic States and Romania. In 2015, wheat was planted on 0.85, 1.97, and 5.11 million acres with 46, 160, and 307 million bushels production in Latvia, Lithuania, and Romania, respectively (USDA FAS, 2016). Tan spot, caused by the fungus Pyrenophora tritici-repentis (Died) Shoem, is an important leaf disease of wheat in the Baltic States (Latvia and Lithuania), Romania, and in many other wheat-growing countries (Bankina and Priekule, 2011; De Wolf et al., 1998; Hosford, 1982; Ronis et al., 2009). The disease was observed in the early 1990s at insignificant level in Baltic States and Romania and since then it is on the rise in the region (Bankina and Priekule, 2011). Increase in tan spot in these countries is thought to be the result of a large acreage of wheat grown under high yielding cultivars but potentially susceptible to tan spot, shift in cultural practices from conventional to minimum tillage that provide ample opportunity for residue borne pathogens like P. tritici-repentis inoculum buildup, and monoculture (Bankina and Priekule, 2011). Tan spot can cause yield losses from 3-50% (Shabeer and Bockus, 1988). In Lithuania, 73% reduction in one thousand kernels weight has been reported depending on the cultivar susceptibility level and inoculum level, and suitable weather conditions for disease development (Bankina and Priekule, 2011; Ronis et al., 2009).
Tan spot can primarily be managed through fungicide application; however, deployment of resistant cultivars seems to be a more economical, environmentally safe and durable approach. Excessive use of fungicide as a disease management strategy could create selection pressure leading to evolution of fungicide resistance in a pathogen population. Fungicide resistance in P. tritici-repentis population has been documented from various countries including Germany and Denmark (Sierotzki et al., 2007).
Development and deployment of durable disease resistant cultivars requires complete information about the pathogen virulence. Variation in P. tritici-repentis virulence has been observed based on using both quantitative, i.e., % necrotic leaf area, lesion size, and lesion number or combination of all the three components (da Luz and Hosford, 1980; Krupinsky, 1992; Misra and Singh, 1972; Schilder and Bergstrom, 1990), and qualitative (i.e., lesion type [Lamari et al., 2003]) rating scales. Lamari and Bernier (1989) for the first time observed two distinct leaf spot symptoms, tan necrosis and chlorosis, produced by P. tritici-repentis on susceptible wheat genotypes. Based on these two distinct symptoms, they developed a pathotype classification system for detecting physiological variation in the fungal population. They grouped the isolates into four pathotypes based on their ability to produce necrosis (nec+) and chlorosis (chl+) and designated them as pathotype 1 (nec+chl+), 2 (nec+chlã), 3 (necãchl+) and 4 (necãchlã) based on their reaction on four bread wheat genotypes, Glenlea, 6B365, 6B662, and Salamouni. Later on, some P. tritici-repentis isolates were found that also produced chlorosis on 6B662 instead 6B365 of the four genotypes and could not be grouped under any of the four identified pathotypes (Lamari et al., 1995). To accommodate these P. tritici-repentis isolates with new virulence, they renamed the pathotype system as race classification system (Lamari et al., 2003). They designated the pathotypes 1-4 as races 1-4 and the newly identified isolates as race 5. The race classification is well-accepted system by the research community working on this pathogen and it has enhanced the researchersã ability to understand wheattan spot host pathogen system.
Till date eight races have been identified in the pathogen populations. Three host-selective toxins Ptr ToxA, Ptr ToxB, and Ptr ToxC have been isolated from the fungal isolates that are associated with necrosis and chlorosis symptoms development and serve as either pathogenicity or aggressiveness factors (Ciuffetti and Tuori, 1999; Friesen et al., 2003). Of these three host-selective toxins, Ptr ToxA and Ptr ToxB are proteinous in nature and have been sequenced and the toxins gene specific primers are available for studying the genotypes of known P. triticirepentis isolates (Andrie et al., 2007). In contrast to these two toxins, Ptr ToxC is a non-ionic, polar, and low molecular weight molecule (Effertz et al., 2002). Ptr ToxC has not been purified, the gene encode it has been not cloned, it is not possible to characterize the fungal isolates for presence of Ptr ToxC gene without testing them on a Ptr ToxC sensitive differential line 6B365.
Several independent studies conducted on the physiological variation in P. tritici-repentis populations have observed diversity in race structure from various countries (Aboudkhaddour et al., 2013; Ali and Francl, 2002; Ali et al., 2004, 2010; Benslimane et al., 2011; Engle et al., 2006; Gamba et al., 2012; Lamari and Bernier, 1989; Lamari et al., 2003; Lepoint et al., 2010; Mikhalova et al., 2014; Moreno et al., 2008; Sarova et al., 2005). Globally race 1 was the most prevalent race (Ali and Francl, 2003; Lamari et al., 2003); however, six races 1, 4, 5, 6, 7, and 8 were reported from Algeria. In USA and Canada races 1-5 have been observed (Ali and Francl, 2002, 2003; Engle et al., 2006; Lamari and Bernier, 1989; Lamari et al., 2003), whereas, races 1 and 2 were observed in South America (Gamba et al., 2012; Moreno et al., 2008).
Since tan spot has been observed in the early 1990s, wheat breeding programs in Baltic States and Romania have developed and released few moderately resistant tan spot cultivars but the disease still impacts crop productivity significantly under high disease pressure (Liatukas et al., 2012). Occurrence of multiple races in the pathogen populations and resistance to individual race controlled by independent genes in the host warrants the investigation of virulence variation in the pathogen populations in the region for development of durable tan spot resistant cultivars. To our knowledge, P. tritici-repentis races prevalent in the Baltic States and Romania on wheat are not available. In this study, we have characterized the fungal isolates from Baltic States and Romania for their race structure using both phenotypic and genotypic approaches.
Two hundred and thirty-one P. triticirepentis isolates from Lithuania (n = 107), Latvia (n = 15), and Romania (n = 109) were used for their race structure in this study (Supplementary Table 1). The isolates were provided by Dr. Z. Liatukas and were received under the Animal and Plant Health Inspection Service (APHIS) permit number P526P-13-02732. The fungal isolates were recovered from tan spot infected leaves samples collected from various wheat growing regions of Lithuania, Latvia, and Romania in 2013 and 2014. Single spore cultures of all 231 isolates were obtained prior to studying their race structure. To obtain single-spore cultures, the isolates were individually grown on V8-PDA medium and conidia were produced as described by Lamari and Bernier (1989). The conidia of each isolate were spread by cutting and rubbing an about 2 cm block of V8-PDA with conidia on fresh V8-PDA plates and single spores were removed with a flamed scalper under stereoscope and grown on a V8-PDA plate. The isolates were stored at ã20ô¯C by following the protocol of Jordahl and Francl (1992) until characterized for their race structure.
DNA of all 231 P. tritici-repentis isolates was obtained by initiating fresh cultures individually on V8-PDA medium by plating frozen dry plugs stored at ã20ô¯C and grew them for 5 days. The mycelia were scraped from the agar surface using a flamed scalpel and placed in a 2 ml microfuge tube. The mycelia was then dried overnight in a water bath at 37ô¯C and then ground into a fine powder using a first prep machine (Retsch MM 301; Glen Mills, Clifton, NJ, USA). DNA was extracted from mycelia of each isolate by following the procedure of Moreno et al. (2008). The DNA concentration normalized to 25 ng/ö¥l using a Nano drop machine (NanoDrop 1000 UV/Vis Spectrophotometer; Counterpane Inc., Tacoma, WA, USA) and run in a 0.8% agarose gel to verify the DNA quality. Genotype of the P. tritici-repentis isolates for Ptr ToxA and Ptr ToxB genes were determined by using the Ptr ToxA and Ptr ToxB genes specific primers developed by Andrie et al. (2007). The conformity of the isolates of P. tritici-repentis was determined by using two P. triticirepentis mating type genes specific primers suggested by Lepoint et al. (2010). PCRs for specific markers were performed in 20 ö¥l volume; 2 ö¥l genomic DNA (25 ng/ö¥l), 0.8 ö¥l of each primer (10 mM), 0.5 ö¥l dNTP (200 ö¥M), 2 ö¥l 10û thermophol buffer, 0.2 ö¥l 10 U/ml Taq Polymerase and 13.7 ö¥l of molecular biology water. PCR reaction was conducted in a S-1000 thermal cycler (BioRad, Hercules, CA, USA) using amplification steps of 94ô¯C for 1 min, followed by 30 cycles of 94ô¯C for 45 s, 55ô¯C for 30 s and 72ô¯C for 1 min with final extension of 72ô¯C for 7 min. The amplified products were electrophoresed on 1.5% agarose gels and scored with reference to 1 kb ladder (New England Biolabs, Beverly, MA, USA). Two known P. tritici-repentis isolates, one each of race 1 and race 5 were used in the experiment as positive checks for Ptr ToxA and Ptr ToxB genes.
Four tan spot wheat differential genotypes Glenlea, 3B365, 6B662, and Salamouni seedlings were raised in 3 û 9 cm plastic containers (Stuewe & Sons Inc., Tangent, OR, USA) filled with Sunshine Mix 1 (Sun Gro Horticulture, Agawam, MA, USA). Three seedlings/container were maintained throughout the experimentation. The plants were watered daily and fertilized once every 10 days post-planting using a slow releasing fertilizer. The plants were kept on a greenhouse bench at 22ô¯C set at 16 h light and 8 h dark cycle. The fungal isolates SD13-101-1 (race 1) and SD13-103 (race 5) recovered from tan spot infected wheat leaves collected in South Dakota were included as positive checks for validation of inoculation and race identification.
Inoculum preparation and inoculations were done as described in Ali and Francl (2001). Briefly, fresh cultures of randomly selected 139 of the 231 isolates genotyped for Ptr ToxA and Ptr ToxB genes were initiated by plating their frozen dry plugs on to fresh V8-PDA medium plates (agar = 10 g; potato dextrose agar [PDA] = 10 g; CaCO3 = 3 g; V8-Juice = 150 ml and 850 distilled water) (Lamari and Bernier, 1989). The plates were incubated in dark for 5 days (when the fungal colony growth generally reaches to 4-5 cm in diameter). The plates were then flooded with distilled sterile water and the hyphal growth was knocked down with the help of a flamed bottom of a test tube. Excess water was removed from the plates prior to incubating them for 24 h in light at room temperature (~22ô¯C) and then at 16ô¯C for 24 h in dark for conidial production. The spores were collected with a flamed looped wire by adding about 30 ml of distilled sterile water in each plate and spore concentration was adjusted at 2,500 spores/ml as described by Jordahl and Francl (1992).
Two-week-old seedlings of all 4 differentials genotypes with three replications were inoculated individually with all 139 isolates by spraying their spore suspension at 2,500 spores/ml using a CO2 pressurized sprayer (www.preval.com) and the inoculated seedlings were placed for 24 h in an automated humidity chamber set at 100% humidity by misting 16 s/10 min for enhancing the chances of fungal infection. Thereafter, the seedlings were moved to greenhouse until rated for symptom development. The isolates were grouped under appropriate race based on their ability to produce necrosis and chlorosis symptoms on appropriate tan spot wheat differentials set developed by Lamari et al. (2003) (Table 1).
P. tritici-repentis mating-type specific DNA bands were amplified in all 231 isolates analyzed conforming the identity of the isolates. As expected, 585 bp and 295 bp DNA bands associated with Ptr ToxA and Ptr ToxB genes, respectively were amplified from the check isolates validating the PCR assay (Fig. 1). Eighty-three percent (n = 191) of the isolates harbored Ptr ToxA gene (Fig. 1, 2, Supplementary Table 2); however, none of the 231 isolates amplified Ptr ToxB gene. Out of the 231 isolates analyzed, 17% (n = 40) did not harbor either of the two toxin genes. Genotypic characterization suggests prevalence (83%) of Ptr ToxA harboring populations (races 1, 2, and 7) in Baltic countries and Romania. The lack of Ptr ToxB gene in the isolates eliminates the possibility of races 5, 6, and 8 in the region as these races contain the ToxB gene (Lamari et al., 2003). The other 17% (n = 40) of the isolates lacked both ToxA and ToxB genes indicating the existence of either race 3 (necãchl+) and/or 4 (necãchlã). Another possibility is that these isolates potentially harbor new virulence genes as previously reported by Ali et al. (2010) and Andrie et al. (2007). They also identified some isolates that behaved like race 1 and race 5 but lacked ToxA and ToxB genes. Due to lack of good molecular markers for Ptr ToxC gene(s), discriminating the fungal isolates that carry Ptr ToxC gene(s) is presently possible by testing isolates on tan spot differential set for their race identification.
A subset of 231 isolates genotyped for Ptr ToxA and Ptr ToxB were evaluated on four differentials to obtain a phenotypic reaction of 139 isolates. A diverse fungal population was prevalent in three countries based on our phenotypic evaluation (Fig. 2). Overall, 63% (n = 88) of the isolates were grouped under race 1 as they produced necrosis and chlorosis on Glenlea and 6B365, respectively and neither symptoms on 6B662 and Salamouni whereas, 8% (n = 11) of the phenotyped isolates were designated as race 2 because they produced necrosis on Glenlea and resistant reaction on the other three differential genotypes. About 11% (n = 15) of the isolates induced chlorosis on 6B365 and neither necrosis or chlorosis on Glenlea, 6B662, and Salamouni and were designated as race 3. Only 5% (n = 7) of the isolates were grouped under race 4, as they did not produce either of the symptoms on all four differential genotypes. Thirteen percent (n = 18) of the isolates could not fit under any of the currently identified 8 races as 17 of them produced necrosis and chlorosis on Glenlea and 6B365 like race 1 but they lacked Ptr ToxA gene. One isolate (RO 5-23) exhibited reactions like race 2 as it produced necrosis on Glenlea and no symptoms on 6B365, 6B662, and Salamouni (Fig. 3), however, lacked Ptr ToxA gene. Thus fungal isolates collected from Lithuania were grouped into races 1, 2, 3, 4, and additional isolates (23%) with different virulence pattern from the currently identified eight races. Race 1 (48%) and race 3 (20%) were the most prevalent races in Lithuania (Fig. 2, Supplementary Table 3) followed by race 4 (8%). Of the fungal isolates from Romania, 80% were grouped as race 1 (n = 43), 17% as race 2 (n = 10), and 2% as race 4 (n = 1). Only one isolate that lacked ToxA gene and behaved like race 2 could not fit under any of the currently identified 8 races. Our results indicate the presence of races 1, 2, and 3 with race 1 the most prevalent race in the region. Presence of races 1-3 on wheat in Balkan state can be expected as all three races with race 1 and 2 at higher frequency were observed in its neighboring country Russia (Mikhalova et al., 2014). The fungal spores can travel long distance through wind (Francl, 1997) and establish far apart from the source of origin under suitable weather conditions. Weather conditions are similar in wheat growing areas on both sides of the border sharing Russia and Baltic State countries. Prevalence of race 1 as a predominant race in the region is not very surprising, this has been observed as the most prevalent race in Algeria (Benslimane et al., 2011), Czech Republic (Ali et al., 2004; Sarova et al., 2005), Canada (Aboudkhaddour et al., 2013; Lamari and Bernier, 1989; Singh and Hughes, 2006), USA (Ali and Francl, 2003; Engle et al., 2006), and South America (Ali and Francl, 2002; Gamba et al., 2012; Moreno et al., 2008). Prevalence of race 1 in higher frequency suggests that wheat germplasm used in the three countries has narrow diversity as far as susceptibility to tan spot races is concerned. Evaluation of both bread wheat and durum wheat genotypes for tan spot using race 1 and race 5 in two independent studies also found higher susceptibility to race 1 as compared to race 5 (Ali et al., 2008; Singh et al., 2006). Prevalence of race 4 in low frequency (5%) in the three countries is likely as this race does not contain any of the three known host-selective toxins and is avirulent on wheat that makes it less competitive with other races for its establishment. Similar trend of prevalence of race 4 at low frequency was reported on wheat from other countries as well (Ali and Francl, 2003; Benslimane et al., 2011; Lamari and Bernier, 1989; Mikhalova et al., 2014; Sarova et al., 2005).
Interestingly, 13% (n = 17) and 2% (n = 1) of the isolates from Lithuania and Romania, respectively did not fit under any of the currently eight identified races though they behave like races 1 and 2. In an earlier study, we reported similar identification of isolates that lack Ptr ToxA gene but are equally virulent like race 1 from Arkansas, USA (Ali et al., 2010). Similarly, 50% of P. tritici-repentis isolates were devoid of Ptr ToxA gene but behaved like race 1 and 2 from Russia, a Balkan state neighboring country (Mironenko et al., 2015). It is likely that wheat cultivars grown in the region, especially in Lithuania may not require Ptr ToxA sensitivity for susceptibility to tan spot. This similar behavior (insensitive to Ptr ToxA and but susceptible to tan spot) has also been reported in wheat cultivars in many independent studies (Ali and Francl, 2002; Ali et al., 2010; Oliver et al., 2008; Noriel et al., 2011). Our results show prevalence of diverse population of P. tritici-repentis, with race 1 (contains Ptr ToxA and Ptr ToxC) being most abundant in the surveyed area. Further, fungal isolates lacking Ptr ToxA gene but behave either like race 1 or 2 were identified suggesting the need further investigate if these isolates produce new toxins or use different arsenals to be pathogenic on wheat. Breeding of tan spot resistant wheat cultivars against race 1 should provide sufficient resistance to minimize losses occurring due to tan spot in the region. However, tan spot resistant germplasm should also be tested against isolates lacking in Ptr ToxA gene but act like races 1 or 2 to be sure if they may carry different toxins from race 1 and hence make race 1 resistant cultivars vulnerable to tan spot. Also, the fungal pathogen should be monitored for any virulence change periodically.
Our results further show that Ptr ToxA and Ptr ToxB genes specific markers can be useful in distinguishing isolates that carry these two genes. However, in the absence of availability of molecular markers for all toxins and the possibility of unknown toxins, characterization of races carrying these toxins by molecular approaches is not realistic. Characterization of P. tritici-repentis races using tan spot wheat differential set for now is the most robust method for exploring variability in the pathogen population until precise molecular markers for all toxins are developed.
Acknowledgments
We like to thank Mr. Richard Gappert for his help in greenhouse and lab work. The South Dakota Wheat Commission and USDA NIFA through SDSU Agricultural Experiment Station (AES) provided financial support for this research.
Fig.ô 1
Gel picture of the Pyrenophora tritici-repentis isolates from Latvia, Lithuania, and Romania showing the presence/absence of ToxA, ToxB and mating type (MT) gene. Lanes 1-3: Ptr ToxA gene in 14-LV-2, 13-LT-5-9, and 14 RO-5-17 isolates (race 1); lanes 4-5: Ptr ToxA gene in 13-LT-1-11 (race 2) and Pti2 of race 1 (positive control); lanes 6-7: isolate from LT as race 3 and race 4; lanes 8-9: necrosis producing isolates 13-LT-2-10 and RO-5-23 without ToxA gene; lanes 10-11: race 5 isolate DW7 (positive control for Ptr ToxB) and water as negative control, respectively.
Fig.ô 2
(A) Genotypic characterization of 223 Pyrenophora tritici-repentis isolates recovered from wheat for Ptr ToxA and Ptr ToxB genes from Latvia, Lithuania, and Romania. The percent of races classified as carrying Ptr ToxA/Ptr ToxB gene or none of them. (B) Race characterization of 139 P. tritici-repentis isolates recovered from wheat collected in Latvia, Lithuania, and Romania. The percent of races classified as race 1, race 2, race 3, race 4 or unknown races. aIsolates lack in Ptr ToxA genes but behave like race 1, bIsolates lack in Ptr ToxA genes but behave like race 2.
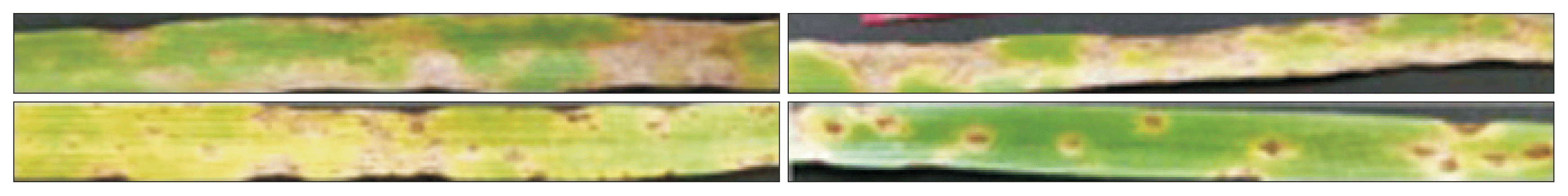
Fig.ô 3
Two Pyrenophora tritici-repentis ToxA- isolates 13-LT-2-10 produced necrosis on Glenlea and chlorosis on 6B365 (left panel top and bottom) and RO-5-23 produced necrosis on Glenlea resistant reaction on 6B365 (right panel top and bottom) behaved like race 1 and 2, respectively.
Tableô 1
Races of Pyrenophra tritici-repentis and their symptoms on tan spot wheat differentials set (Lamari et al., 2003)
References
Aboudkhaddour, R, Turkington, TK and Strelkov, SE 2013. Race structure of Pyrenophora triciti-repentis (tan spot of wheat) in Alberta, Canada. Can J Plant Pathol. 35:256-268.

Ali, S and Francl, LJ 2001. Recovery of Pyrenophora triticirepentis from barley and reaction of 12 cultivars to five races and two host-specific toxins. Plant Dis. 85:580-584.


Ali, S and Francl, LJ 2002. Race structure of Pyrenophora tritici-repentis isolates obtained from wheat in South America. Plant Protect Sci. 38(Special Issue 2):302-304.

Ali, S and Francl, LJ 2003. Population race structure of Pyrenophora tritici-repentis prevalent on wheat and noncereal grasses in the Great Plains. Plant Dis. 87:418-422.


Ali, S, Gurung, S and Adhikari, TB 2010. Identification and characterization of novel isolates of Pyrenophora triticirepentis from Arkansas. Plant Dis. 94:229-235.


Ali, S, Sarova, J and Waukulinsky, W 2004 Pyrenophora tritici-repentis races in the Czech Republic and Poland. Phytopathology. 93(Suppl):S3.(Abstract).
Ali, S, Singh, PK, McMullen, MP, Mergoum, M and Adhikari, TB 2008. Resistance to multiple leaf spot diseases in wheat. Euphytica. 159:167-179.


Andrie, RM, Pandelova, I and Ciuffetti, LM 2007. A combination of phenotypic and genotypic characterization strengthens Pyrenophora tritici-repentis race identification. Phytopathology. 97:694-701.


Bankina, B and Priekule, I 2011. A review of tan spot research in Baltic countries: occurrence, biology and possibilities of control. é§emdirbystá (Agriculture). 98:3-10.
Benslimane, H, Lamari, L, Benbelkacem, A, Sayoud, R and Bouznad, Z 2011. Distribution of races of Pyrenophora tritici-repentis in Algeria and identification of a new virulence type. Phytopathol Mediterr. 50:203-211.
Ciuffetti, LM and Tuori, RP 1999. Advances in the characterization of the Pyrenophora tritici-repentis: wheat interaction. Phytopathology. 89:444-449.


da Luz, WC and Hosford, RM Jr 1980. Twelve Pyrenophora trichostoma races for virulence to wheat in the central plains of North America. Phytopathology. 70:1193-1196.

De Wolf, ED, Effertz, RJ, Ali, S and Francl, LJ 1998. Vistas of tan spot research. Can J Plant Pathol. 20:349-370.

Effertz, RJ, Meinhardt, SW, Anderson, JA, Jordahl, JG and Francl, LJ 2002. Identification of a chlorosis-inducing toxin from Pyrenophora tritici-repentis and the chromosomal location of an insensitivity locus in wheat. Phytopathology. 92:527-533.


Engle, JS, Madden, LV and Lipps, PE 2006. Distribution and pathogenic characterization of Pyrenophora tritici-repentis and Stagonospora nodorum in Ohio. Phytopathology. 96:1355-1362.


Francl, LJ 1997. Local and mesoscale dispersal of Pyrenophora tritici-repentis conidia. Can J Plant Pathol. 19:247-255.
Friesen, TL, Ali, S, Kianian, S, Francl, LJ and Rasmussen, JB 2003. Role of host sensitivity to Ptr ToxA in development of tan spot of wheat. Phytopathology. 93:397-401.


Gamba, FM, Strelkov, SE and Lamari, L 2012. Virulence of Pyrenophora tritici-repentis in the southern cone region of South America. Can J Plant Pathol. 34:545-550.

Hosford, RM Jr 1982. Tan spot: developing knowledge 1902-1981, virulent races and wheat differentials, methodology, rating systems, other leaf diseases, literature. In: Tan spot of wheat and related disease workshop, eds. by RM Hosford, 1-20. North Dakota State University, Fargo, ND, USA.
Jordahl, JG and Francl, LJ 1992. Increase and storage of cultures of Pyrenophora tritici-repentis. In: Advances in tan spot research, eds. by LJ Francl, JM Krupinsky and MP McMullen, 109. North Dakota State University, Fargo, ND, USA.
Lamari, L and Bernier, CC 1989. Evaluation of wheat lines and cultivars to tan spot (Pyrenophora tritici-repentis) based on lesion type. Can J Plant Pathol. 11:49-56.

Lamari, L, Sayoud, R, Boulif, M and Bernier, CC 1995. Identification a new race in Pyrenophora tritici-repentis: implications for the current pathotype classification system. Can J Plant Pathol. 17:312-318.
Lamari, L, Strelkov, SE, Yahyaoui, A, Orabi, J and Smith, RB 2003. The identification of two new races of Pyrenophora tritici-repentis from the host center of diversity confirms a one-to-one relationship in tan spot of wheat. Phytopathology. 93:391-396.


Lepoint, P, Renard, ME, Legrû´ve, A, Duveiller, E and Maraite, H 2010. Genetic diversity of the mating type and toxin production genes in Pyrenophora tritici-repentis. Phytopathology. 100:474-483.


Liatukas, Z, Ruzgas, V, Razbadauskiená, K, Brazauskas, G and Koppel, R 2012. Winter wheat cultivars ãKovas DSã, ãZunda DSã, ãVikaras DSã, ãKaskada DSã for high input farming: development and characterization. é§emdirbystá (Agriculture). 99:255-264.
Mikhalova, LA, Mironenko, NV and Kovalenko, NM 2014. Populations of Pyrenophora tritici-repentis in the North Caucasus and North-West Russia: the racial composition and dynamics of virulence. Mikol Fitopatol. 48:393-400.
Mironenko, NV, Baranova, OA, Kovalenko, NM and Mikhalova, LA 2015. Frequency of ToxA gene in North Caucasian and North-West Russian populations of Pyrenophora tritici-repentis. Mikol Fitopatol. 49:325-329.
Misra, AP and Singh, RA 1972. Pathogenic differences amongst three isolates of Helminthosporium tritici-repentis and the performance of wheat varieties against them. Indian Phytopathol. 25:350-353.
Moreno, MV, Stenglein, SA, Balatti, PA and Perellû°, AE 2008. Pathogenic and molecular variability among isolates of Pyrenophora tritici-repentis, causal agent of tan spot of wheat in Argentina. Eur J Plant Pathol. 122:239-252.


Noriel, AJ, Xiaochun, S, Willium, B and Guihua, B 2011. Resistance to tan spot and insensitivity to Ptr ToxA in wheat. Crop Sci. 51:1059-1067.


Oliver, RP, Lord, M, Rybak, K, Faris, JD and Solomon, PS 2008. Emergence of tan spot disease caused by toxigenic Pyrenophora tritici-repentis in Australia is not associated with increased deployment of toxin-sensitive cultivars. Phytopathology. 98:488-491.


Ronis, A, SemaéÀkiene, R, Dabkeviáius, Z and Liatukas, é§ 2009. Influence of leaf diseases on grain yield and yield components in winter wheat. J Plant Prot Res. 49:151-157.

Sarova, J, HanzalovûÀ, A and BartoéÀ, P 2005. Races of Pyrenophora tritici-repentis in Czech Republic. Acta Agrobot. 58:73-78.


Schilder, AMC and Bergstrom, GC 1990. Variation in virulence within the population of Pyrenophora tritici-repentis in New York. Phytopathology. 80:84-90.

Shabeer, A and Bockus, WW 1988. Tan spot effects on yield and yield components relative to growth stage in winter wheat. Plant Dis. 72:599-602.

Sierotzki, H, Frey, R, Wullschleger, J, Palermo, S, Karlin, S, Godwin, J and Gisi, U 2007. Cytochrome b gene sequence and structure of Pyrenophora teres and P. tritici-repentis and implications for QoI resistance. Pest Manag Sci. 63:225-233.


Singh, PK and Hughes, GR 2006. Genetic similarity among isolates of Pyrenophora tritici-repentis, causal agent of tan spot of wheat. J Phytopathol. 154:178-184.

Singh, PK, Mergoum, M, Ali, S, Adhikari, TB, Elias, EM, Anderson, JA, Glover, KD and Berzonsky, WA 2006. Evaluation of elite wheat germ plasm for resistance to tan spot. Plant Dis. 90:1320-1325.


[USDA FAS] USDA Foreign Agricultural Service. 2016 onwards Gain report. URL http://gain.fas.usda.gov/Recent%20GAIN%20Publications/Grain%20and%20Feed%20Annual%20-%20Lithuania%202016_Warsaw_Lithuania_4-14-2016.pdf. 4 January 2016.
- TOOLS
-
METRICS

- Related articles
-
Characterization of
Melon necrotic spot virus Occurring on Watermelon in Korea2015 December;31(4)








 PDF Links
PDF Links PubReader
PubReader Full text via DOI
Full text via DOI Full text via PMC
Full text via PMC Download Citation
Download Citation Supplement
Supplement Print
Print