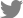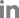|
 |
| Plant Pathol J > Volume 33(6); 2017 > Article |
Abstract
The clpks18 gene was first cloned and identified in Curvularia lunata. It contains 6571 base pairs (bp) and an 6276 bp open reading frame encoding 2091 amino acids. The ClPKS18 deletion mutant displayed an albino phenotype, and almost lost the ability to product 5-(hydroxymethyl) furan-2-carboxylate (M5HF2C) toxin, implying that clpks18 gene in C. lunata is not only involved in 1,8-dihydroxynaphthalene melanin synthesis, but also relatively associated with M5HF2C toxin biosynthesis of the pathogen. The pathogenicity assays revealed that ΔClPKS18 was impaired in colonizing the maize leaves, which corresponds to the finding that ClPKS18 controls the production of melanin and M5HF2C in C. lunata. Results indicate that ClPKS18 plays a vital role in regulating pathogenicity of in C. lunata.
Curvularia lunata is an important fungal pathogen that causes Curvularia leaf spot (CLS), which is one of the most widely distributed maize leaf diseases worldwide (Gao et al., 2015a; Liu et al., 2016). To uncover the pathogenicity mechanisms of CLS, most researches have focused on identifying virulence factors, such as melanin (Gao et al., 2012, 2015b), and toxin (Gao et al., 2014a; Liu et al., 2009). Fungal melanins, formed by the oxidative polymerisation of phenolic compounds, are essential for enhance the mechanical strength of the infection into the host plant epidermis and contribute to virulence in numerous types of plant diseases (Gao et al., 2015e). Previous studies have also indicated that the albino C. lunata strains present in wild-type (WT) strains are weakly virulent to maize leaves, supporting the conclusion that melanin is a crucial virulence factor for C. lunata in susceptible maize leaves (Yan et al., 2005). Researchers have successfully identified a non-host-specific toxin called methyl 5-(hydroxymethyl) furan-2-carboxylate (M5HF2C). The super-virulence capability of M5HF2C was also observed on some other plant species, such as Capsicum annuum, Nicotiana tabacum, Oryzae sativa, and (Liu et al., 2009). However, few genes involved in the production of melanin (Gao et al., 2015d) and toxin (Gao et al., 2015c) have been researched.
The PKS18 protein is involved in melanin synthesis in some pathogens, such as Cochliobolus heterostrophus (Eliahu et al., 2007). However, no research has revealed the exact correlation of PKS18 with melanin production and pathogenicity in C. lunata. This study first reports the sequence and characterization of pks18 gene from C. lunata. The Bipolaris maydis PKS18 (accession number: AY495659) was used to query the C. lunata genome database (Dryad Digital Repository) for orthologs (Gao et al., 2014b). The open reading frame of clpks18 comprises 6571 bp and an 6276 bp open reading frame encoding a 209-amino-acid protein. Phylogenetic analysis indicated that the ClPKS18 protein falls in a well-supported group of dothideomycete polyketide synthetase homologs (Fig. 1A). Alignment of ClPKS18 with B. maydis BmPKS18 and Setosphaeria turcica StPKS18 showed 88% identity and 99% positives, respectively (NCBI BlastP Align). Meanwhile, alignment of them identified highly conserved Acyl transferase, Beta-ketoacyl synthase, malonyl CoA-acyl carrier protein transacylase, and dehydratase domains, characteristic of fungal polyketide synthases (Fig. 1B).
Target gene deletion strategy was employed by replacing clpks18 with a hygromycin resistance (hph) cassette to investigate the biological functions of ClPKS18 in C. lunata (Fig. 2A). The Southern hybridization pattern confirmed that homologous recombination occurs at the clpks18 locus in ΔClPKS18 (Fig. 2B). The radial growth rates of the mutant and WT on the potato dextrose agar (PDA) medium were compared. ΔClPKS18 had a significantly slower mycelial growth rate than WT on the PDA medium. However, the conidiation of ΔClPKS18 was not evidently different from those of the WT (Table 2).
In B. maydis, BmPKS18 was functionally characterized to encode a polyketide synthetase that converts malonyl-CoA to 1,3,6,8-tetradroxynaphthalene (1,3,6,8-THN) in the DHN-melanin synthesis pathway (Eliahu et al., 2007). This research showed that the ClPKS18 deletion mutant of C. lunata displayed an albino phenotype, indicating that ClPKS18 is an orthologue of PKS18 from B. maydis (Fig. 3A). We detected the expression of the polyketide synthase gene (pks18), the transcription factor gene (cmr1), and three synthase genes (brn1, brn2, and scd) related to the synthesis of DHN melanin in the mutant and WT to further confirm this observation (Gao et al., 2017). As expected, almost no expression of pks18 was detected in ΔClPKS18. Besides, the expression of scd in ΔClPKS18 has a 70.92-fold decrease, and the expression of brn1 and brn2 have above 30-fold decrease in ΔClPKS18 compared to those in WT. As for cmr1, it also showed a downward trend, which is almost 0.36-fold decrease in ΔClPKS18 compared with WT (Fig. 3B). Overall, we conclude that ClPKS18 plays a positive regulation role in the synthesis of melanin.
The mutants were cultured in Fries 3 medium for 30 days to determine whether they retained the ability to produce the virulence-related toxin M5HF2C (Liu et al., 2009). As shown in Fig. 4, the mutant lost the ability to produce M5HF2C toxin. The expression of the M5HF2C biosynthesis related gene clt-1 was analyzed by qRT-PCR to further confirm that ΔClPKS18 acts as a positive regulator of M5HF2C toxin production. Meanwhile, the expression of brn1, which is also responsible for M5HF2C biosynthesis (Liu et al., 2011), fell sharply in ΔClPKS18 (Fig. 3B). The experiment results indicate that ClPKS18 played a major role in the regulation of M5HF2C biosynthesis in C. lunata. We further assayed the infective ability of ΔClPKS18 on maize leaves because the deletion of clpks18 compromised the ability of C. lunata to produce M5HF2C. Similarly, ΔClPKS18 lost the pathogenicity, and the lesion areas also cannot be found in maize leaves which treated with ΔClPKS18 (Fig. 5), indicating that ClPKS18 was essential to the complete virulence in C. lunata. We proposed that the clpks18 gene may be one of the node genes controlling two separate metabolic pathways for melanin and toxin biosynthesis, respectively. In conclusion, this study would help us understand the synthesis mechanism of melanin and toxin in C. lunata and may provide target sites for designing a new agent to control C. lunata and a few similar fungi.
Acknowledgments
We would like to thank Zhe Li for critical reading of the manuscript. The work was supported by the National Science Foundation of China (31471734) and China Agriculture Research System (CARS-02).
Fig.┬Ā1
C. lunata PKS18 is an ortholog of Biolaris maydis PKS18. (A) Phylogenetic analysis. PKS18 protein sequences were obtained from GenBank using B. maydis BmPKS18 as a query. (B) C. lunata ClPKS18, B. maydis BmPKS18 and Setosphaeria turcica StPKS18 were aligned using ClustalW. Conserved Acyl transferase domains are highlighted in blue, Beta-ketoacyl synthase domains are highlighted in yellow, malonyl CoA-acyl carrier protein transacylase domains are highlighted in red, dehydratase domains are highlighted in green, asterisks mark identical residues, colons mark conserved residues, and periods indicate semi-conserved residues.
GenBank Accession numbers
Dothideomycetes: Alternaria alternate AaPKS18: AFN68292; Bipolaris maydis BmPKS18: AAR90272; Curvularia lunata ClPKS18: MF114294; Dothistroma septosporum DsPKS18: EME39782; Pyrenophora tritici-repentis PtPKS18: XP_001933656; Setosphaeria turcica StPKS18: AEE68981.
Sordariomycetes: Verticillium dahlia VdPKS18: AGI15329; Verticillium longisporum VlPKS18: CRK15634.
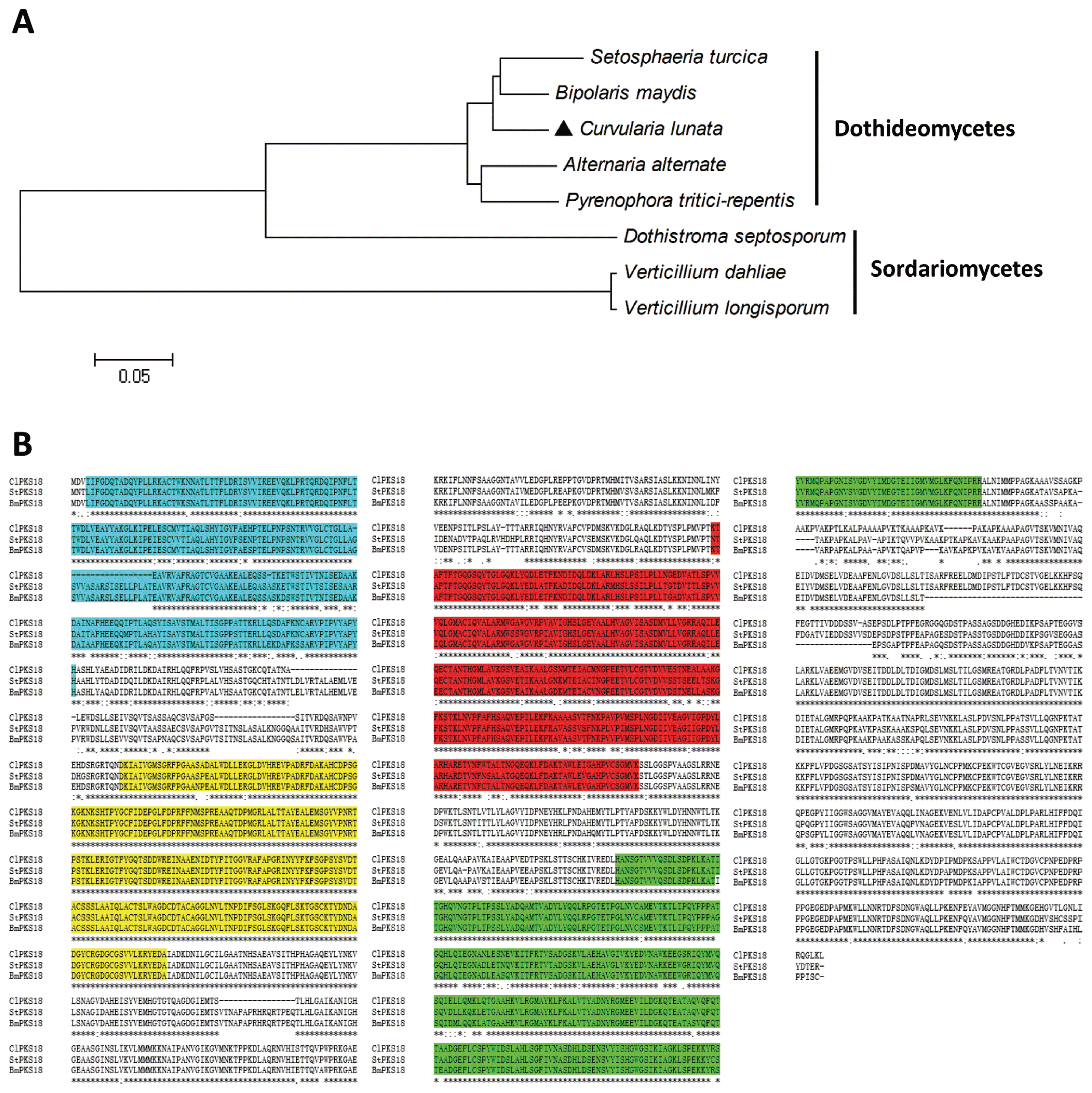
Fig.┬Ā2
Graphical representation of screening clpks18 deletion mutant. (A) clpks18 deletion strategy used by homologous recombination. clpks18 and hygromycin resistance (hph) genes are represented by green and red boxes, respectively. (B) Southern blot to confirm clpks18 was replaced using the 600 bp fragment of clpks18 as probe. All primers used for gene deletion and confirmation are showed in Table 1.

Fig.┬Ā3
ClPKS18 possively regulates the mycelial melanization of C. lunata. (A) Cultures of WT strain (CX-3), and clpks18 deletion mutant (ΔClPKS18) grown on PDA plates. Note the white mycelia of ΔClPKS18 compared to WT. (B) qRT-PCR analyses of pks18, cmr1, brn1, brn2, and scd. Error bars are the standard deviation. A single asterisk indicates the P < 0.05 while double asterisks indicate the P < 0.001 in the T-test analysis. All primers used for qRT-PCR are showed in Table 1.

Fig.┬Ā4
ClPKS18 regulates the biosynthesis of M5HF2C toxin. HPLC-MS chromatograms of the methyl 5-(hydroxymethyl)-furan-2-carboxylate standard and toxins extracted from the WT strain (CX-3) and clpks18 deletion mutant (ΔClPKS18).
Fig.┬Ā5
Virulence of the WT (CX-3), and clpks18 deletion mutant (ΔClPKS18) on maize leaves. ΔClPKS18 is impaired in the colonization of maize leaves. Detached leaves of HUANGZAO-4 were inoculated with conidial suspensions and incubated on two layers of filter papers moisturized with 10 mM 6-Benzyladenine (6-BA) in Petri dishes at 28°C for 96 h.

Table┬Ā1
Primer used for this study
| Primers | Sequence 5ŌĆ▓ to 3ŌĆ▓ | Description | PCR (kb)a | Purpose |
|---|---|---|---|---|
| pks18-FL-F | ATGGATGTCATCATCTTCGG | clpks18 coding region, FPb | 6.571 | dclpks18 confirmation |
| pks18-FL-R | TTATAGCTTAAGGCCTTGCC | clpks18 coding region, RPc | ||
|
|
||||
| pks18U-F | ACGCGTCGACACCCGCAGGATATATCA | flanking region upstream of clpks18 FP | 0.925 | ΔClPKS18 |
| pks18U-R | GCTCTAGATTTAGCAAGGGTATAATT | flanking region upstream of clpks18 RP | ||
|
|
||||
| pks18D-F | GGGGTACCGCATTTTGATTATTCATC | flanking region downstream of clvelB FP | 0.943 | |
| pks18D-R | CGGAATTCCCAGAGGCATGGGTCGGT | flanking region downstream of clvelB RP | ||
|
|
||||
| hyg-F | CGACAGCGTCTCCGACCTGA | hph coding region, FP | 0.811 | ΔClPKS18 confirmation |
| hyg-R | CGCCCAAGCTGCATCATCGAA | hph coding region, RP | ||
|
|
||||
| pks18-sb-F | AGATTGTCTCTCAGGTCACC | clpks18 coding region, FPb | 0.600 | |
| pks18-sb-R | AATTGATTCTGCCAGGCGCG | clpks18 coding region, RPc | ||
|
|
||||
| gapdh_2F | TCGTCGCCGTAAACGACCCC | gapdh coding region, FP | 0.207 | qRT-PCR |
| gapdh_2R | CGCCCTTGAACTGGCCGTGT | gapdh coding region, RP | ||
|
|
||||
| brn1-1F | TGGCCAGCCAGTAGACATTG | brn1 coding region, FP | 0.075 | |
| brn1-1R | ACCTTTCCGTTGACCCACTC | brn1 coding region, RP | ||
|
|
||||
| brn2-2F | AACAACGGCCGTATCATCCT | brn2 coding region, FP | 0.078 | |
| brn2-2R | AGCGTTGTAAAGAGCGTGGT | brn2 coding region, RP | ||
|
|
||||
| cmr1-1F | GTTTGGACTGACTCGCTGGT | cmr1 coding region, FP | 0.118 | |
| cmr1-1R | TAGGATGATCGGCGGGAAGA | cmr1 coding region, RP | ||
|
|
||||
| scd_1F | CGGTCGTTCCTGGACAAGATG | scd coding region, FP | 0.121 | |
| scd_1R | GTGTGCCTCCGATAAAGTGCTG | scd coding region, RP | ||
|
|
||||
| clt1-F | GCACACACATACCCAAGACG | clt-1 coding region, FP | 0.150 | |
| clt1-R | AGTTGATGGGAATGTAGGCG | clt-1 coding region, RP | ||
References
Eliahu, N, Igbaria, A, Rose, MS, Horwitz, BA and Lev, S 2007. Melanin biosynthesis in the maize pathogen Cochliobolus heterostrophus depends on two mitogen-activated protein kinases, Chk1 and Mps1, and the transcription factor Cmr1. Eukaryot Cell. 6:421-429.



Gao, JX, Gao, SG, Li, YQ and Chen, J 2015a. Genome-wide prediction and analysis of the classical secreted proteins of Curvularia lunata. J Plant Prot. 42:869-876.
Gao, JX, Jing, J and Chen, J 2015b. Elementary coordinated expression research on genes related to the synthesis of pathogenesis-related melanin and toxin in Cochliobolus lunatus. J SJTU Agr Sci. 33:53-58.
Gao, JX, Jing, J, Liu, T and Chen, J 2015c. Identification of Clt-1-regulated proteins associated with the production of non-host-specific toxin and pathogenicity in Cochliobolus lunatus. J Phytopathol. 163:33-41.

Gao, JX, Jing, J, Liu, T, Yu, CJ, Li, YQ and Chen, J 2015d. Identification of proteins associated with the production of melanin and with pathogenicity in maize pathogen Curvularia lunata. Australas Plant Pathol. 44:599-603.

Gao, JX, Jing, J, Yu, CJ and Chen, J 2015e. Construction of a high-quality yeast two-hybrid library and its application in identification of interacting proteins with Brn1 in Curvularia lunata. Plant Pathol J. 31:108-114.


Gao, JX, Liu, T and Chen, J 2014a. Insertional mutagenesis and cloning of the gene required for the biosynthesis of the non-host-specific toxin in Cochliobolus lunatus that causes maize leaf spot. Phytopathology. 104:332-339.


Gao, JX, Yu, CJ, Wang, M, Sun, JN, Li, YQ and Chen, J 2017. Involvement of a velvet protein ClVelB in the regulation of vegetative differentiation, oxidative stress response, secondary metabolism, and virulence in Curvularia lunata. Sci Rep. 7:46054



Gao, SG, Li, YQ, Gao, JX, Suo, YJ, Fu, KH, Li, YY and Chen, J 2014b. Genome sequence and virulence variation-related transcriptome profiles of Curvularia lunata, an important maize pathogenic fungus. BMC Genomics. 15:627


Gao, SG, Liu, T, Li, YY, Wu, Q, Fu, KH and Chen, J 2012. Understanding resistant germplasm-induced virulence variation through analysis of proteomics and suppression subtractive hybridization in a maize pathogen Curvularia lunata. Proteomics. 12:3524-3535.


Liu, T, Liu, LX, Jiang, X, Huang, XL and Chen, J 2009. A new furanoid toxin produced by Curvularia lunata, the causal agent of maize Curvularia leaf spot. Can J Plant Pathol. 31:22-27.

Liu, T, Wang, YY, Ma, BC, Hou, JM, Jin, YZ, Zhang, YL, Ke, XW, Tai, LM, Zuo, YH and Dey, K 2016. Clg2p interacts with Clf and ClUrase to regulate appressorium formation, pathogenicity and conidial morphology in Curvularia lunata. Sci Rep. 6:24047



Liu, T, Xu, SF, Liu, LX, Zhou, FH, Hou, JM and Chen, J 2011. Cloning and characteristics of Brn1 gene in Curvularia lunata causing leaf spot in maize. Eur J Plant Pathol. 131:211-219.

Yan, H, Chen, J, Gao, Z, Xia, S and Zhang, R 2005. The heredity and variation of the interaction between Curvularia Lunata and Plant. J Maize Sci. 13:119-120.
- TOOLS



 PDF Links
PDF Links PubReader
PubReader Full text via DOI
Full text via DOI Full text via PMC
Full text via PMC Download Citation
Download Citation Print
Print