Reverse Transcription Recombinase Polymerase Amplification Assay for Rapid and Sensitive Detection of Barley Yellow Dwarf Virus in Oat
Article information
Abstract
Barley yellow dwarf virus (BYDV) is an economically important plant pathogen that causes stunted growth, delayed heading, leaf yellowing, and purple leaf tip, thereby reducing the yields of cereal crops worldwide. In the present study, a reverse transcription recombinase polymerase amplification (RT-RPA) assay was developed for the detection of BYDV in oat leaf samples. The RT-RPA assay involved incubation at an isothermal temperature (42°C) and could be performed rapidly in 5 min. In addition, no cross-reactivity was observed to occur with other cereal-infecting viruses, and the method was 100 times more sensitive than conventional reverse transcription polymerase chain reaction. Furthermore, the assay was validated for the detection of BYDV in both field-collected oat leaves and viruliferous aphids. Thus, the RT-RPA assay developed in the present study represents a simple, rapid, sensitive, and reliable method for detecting BYDV in oats.
In comparison to other cereal species, oat contains high levels of compounds that exert sedative and soothing effects on the brain and nervous system as well as keep blood fat under control (Peterson and Wood, 1997). Spurred by increasing interest in healthy foods, oat production in Korea has recently increased.
Barley yellow dwarf virus (BYDV), which is a member of the genus Luteovirus (Luteoviridae), is one of the most common pathogens of cereal crops throughout the world (Jedlinski, 1981). BYDV-infected oat plants exhibit yellowing or reddening, stunted growth, an upright posture of thickened stiff leaves, and reduced yield (McKirdy et al., 2002). As with other luteoviruses, BYDV can be transmitted by a variety of aphid vectors (Rochow and Duffus, 1981), such as Rhopalosiphum padi and Sitobian avanae, between the phloem of infected and un-infected plants and can cause physiological, biochemical, and cytological alteration, including the degeneration of phloem and restriction of photosynthate transportation. BYDV-infected plants are also more prone to other pathogens and abiotic stresses (Scheller and Shukle, 1986). Previous studies have reported that the incidence, pattern, severity, rate, and timing of infection by aphids are influenced by environmental conditions (Fabre et al., 2005). In particular, humid and mild climates affect the continuity development and reproduction of aphids (Roos et al., 2011), and the incidence of BYDV generally increases with increases in the size of aphid populations. Thus, rapid diagnostic detection is critical to preventing the spread of BYDV.
Several methods have been used to detect BYDV in cereal crops, and these include reverse transcription polymerase chain reaction (RT-PCR), multiplex RT-PCR, and real-time TaqMan RT-PCR (Balaji et al., 2003; Canning et al., 1996). However, these methods are time-consuming, laborious, expensive, require trained personnel, and must generally be performed in a laboratory setting (Martínez-Culebras et al., 2001; Wang et al., 2009). As an alternative, loop-mediated isothermal amplification (LAMP) assays have been developed to overcome the disadvantages associated with PCR and RT-PCR (Zhao et al., 2010). However, RT-LAMP assays require four or six primers that are difficult and, thus, expensive to design and require the use of relatively high temperatures, which make such assays difficult and impractical for use in the field.
Another alternative to PCR and RT-PCR methods is recombinase polymerase amplification (RPA), which is an isothermal nucleic acid amplification technique that is generally considered simple, rapid, sensitive, and costeffective (Piepenburg et al., 2006). Indeed, RPA reactions can be performed at low temperatures (37-42°C) and, under ideal conditions, can be completed in under 40 min. For RNA targets, such as BYDV, a reverse transcription step is required to synthesize cDNA prior to the RPA reactions (Euler et al., 2012). However, the resulting RT-RPA amplification can then be easily visualized using agarose gel electrophoresis. To promote the use of RT-RPA for the detection of BYDV in oat crops, the aim of the present study was to compare the performances of RT-RPA and the more widely known RT-PCR and to develop a rapid, sensitive, and reliable RT-RPA assay for detecting BYDV in both oats and aphids.
In the present study, BYDV-infected oat leaves were collected from oat farms located in Kangjin and Jeongeup, Korea, in April 2020, and aphids (R. padi) were collected from oat leaves that exhibited BYDV disease symptoms. Total RNA was extracted from the samples using the Clear-S Total RNA Extraction Kit (InVirusTech Co., Gwangju, Korea), according to the manufacturer’s instructions. Most of the samples were screened for BYDV using RT-PCR, previously described diagnostic primers (Malmstrom and Shu, 2004), the SuPrimeScript RT-PCR Premix (GeNet Bio, Daejeon, Korea), according to the manufacturer’s instructions, and the following reaction conditions: 50°C for 30 min; 95°C for 5 min; 35 cycles of 95°C for 30 s, 56°C for 60 s, and 72°C for 60 s; and 72°C for 10 min. After amplification, the resulting RT-PCR products were visualized by gel electrophoresis using 2% agarose gels.
To design optimal primers for the BYDV RT-RPA assay, 25 complete BYDV coat protein (CP) gene sequences (LC530629, AF235167, EF521846, DQ118372, EF521840, KY593458, X56050, AY879231, KY593457, DQ118372, X07653, MK962883, MK012661, MK012655, KT252975, KT252976, KP771878, EU332333, EU332332, AY855920, MF693133, MF693131, KU097016, MF693124, and LC530629) were obtained from the GenBank database and aligned, for identification of highly conserved regions, using BioEdit version 7.0.5.3. Three sets of primers were then selected using PrimedRPA (TwistDx, Ltd., Cambridge, UK), according to the manufacturer’s instructions (Table 1), and then evaluated to select the best primer set. The BYDV182 primer set, which amplified a product of 182 bp, was selected for the assay since the other primer sets did not amplify the expected amplicons or increase non-specific bands (data not shown). The RT-RPA products of the complete BYDV CP were determined by cloning the target into the pGEM-T Easy Vector (Promega, Madison, WI, USA) and then sequencing the cloned fragments.
Conventional RT-RPA assays were performed using the TwistAmp Basic RT kit (TwistDx, Ltd.). Briefly, each 50 μl reaction contained 0.5 μg total RNA, 0.48 μM of each primer, 29.5 μl 1× rehydration buffer, and 2.5 μl magnesium acetate (280 nM). cDNA was then synthesized by incubating the resulting products with 1 μl SuperScript III (Invitrogen, Carlsbad, CA, USA) and 1.2 μl RNase inhibitor in a 42°C water bath for 20 min and visualized using gel electrophoresis on 2% agarose gels for 40 min. Total RNA from confirmed BYDV-infected oat samples were used as positive controls for both the RT-RPA and RT-PCR assays, and total RNA from BYDV-free samples were used as negative controls.
To optimize the reaction time of the RT-RPA assay, the effectiveness of 1, 5, 10, 15, 20, and 30 min reaction times were compared. Only the 5-min reaction time yielded clear amplicon with the expected size (182 bp), and no significant difference was observed in the product yields when the reaction time was increased further (Fig. 1). Three independent reaction times for optimization experiments using different RNA samples were set, which yielded similar results. This result suggested that a 5-min incubation was sufficient for the amplification of BYDV.
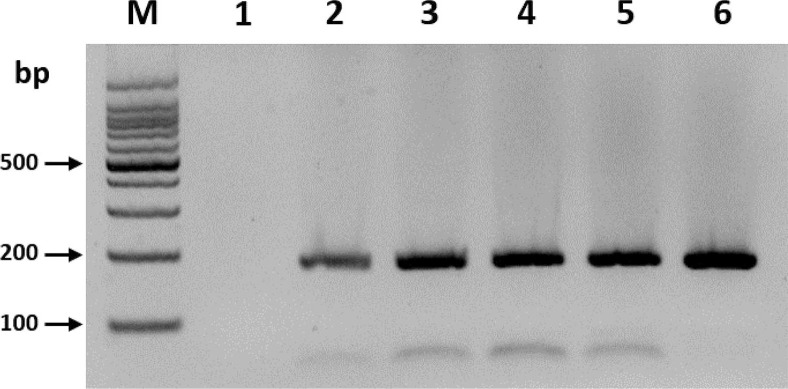
Optimization of reverse transcription recombinase polymerase amplification reaction time on the detection of barley yellow dwarf virus (BYDV). Lane M, DNA marker; lanes 1-6, reaction times of 1, 5, 10, 15, 20, and 30 min, respectively.
To compare the sensitivities of the RT-RPA and RT-PCR assays, the complete BYDV CP transcripts obtained from cloned plasmids using T7 RNA polymerase (Promega) were used for serial dilutions. The synthesized transcript concentrations were measured using a Nanodrop Spectrophotometer (BioDrop, Cambridge, UK), and a ten-fold serial dilution series (100-10-7) of BYDV CP transcripts (~500 ng/μl) was prepared using RNase-free water. Visualization of the resulting products on 2% agarose gels revealed that the RT-RPA assay consistently detected even the smallest amount of BYDV CP transcripts, i.e., 50 fg/μl RNA (10-7 dilution), whereas the RT-PCR assay was only able to detect BYDV CP transcripts as low as 5 pg/μl RNA (10-5 dilution) (Fig. 2). Thus, the RT-RPA assay was at least 100 times more sensitive than RT-PCR for the detection of BYDV. Three independent reactions were carried out, and similar results were observed.
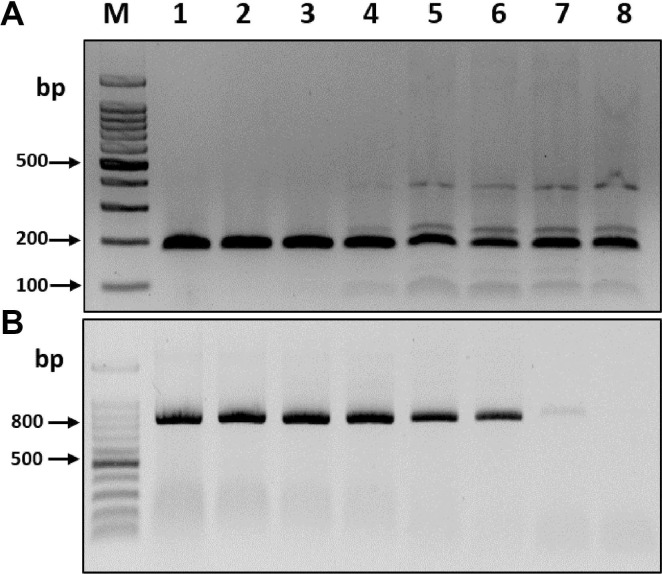
Comparison of the amplification method on the sensitivity of barley yellow dwarf virus (BYDV) detection. (A) Reverse transcription recombinase polymerase amplification (RT-RPA). (B) Reverse transcription polymerase chain reaction (RT-PCR). Amplified products (182 bp from the RT-RPA assay and 831 bp from RT-PCR) were visualized on 1.2% agarose gels. M, DNA marker; lanes 1-8, 10-fold serial dilution (100-10-7) of BYDV template (coat protein transcripts).
To compare the specificities of the RT-RPA and RT-PCR assays, the leaves of both healthy oat plants and those infected with other major cereal-infecting viruses, including barley mild mosaic virus (BaMMV) and barley yellow mosaic virus (BaYMV), were screened using conventional RT-PCR assay for BaMMV and BaYMV and previously published specific primers (Lee et al., 2017). The RT-PCR products were analyzed using 2% agarose gel electrophoresis. The RT-RPA assay did not amplify either BaMMV or BaYMV, which demonstrated the high specificity of the method (Fig. 3).
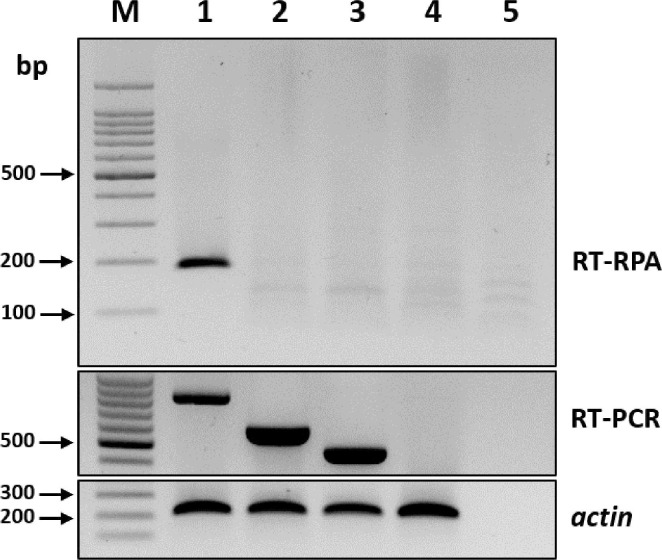
Specificity of the reverse transcription recombinase polymerase amplification (RT-RPA) assay for the detection of barley yellow dwarf virus (BYDV). RT-PCR, reverse transcription polymerase chain reaction; M, DNA marker; lanes 1-4, RT-RPA product amplified from BYDV-, barley mild mosaic virus (BaMMV), and barley yellow mosaic virus (BaYMV)-infected (lanes 1-3) and healthy (lane 4) tissues, respectively; lane 5, negative control. An actin was used as an internal control.
To assess the reliability of RT-RPA assays for detecting BYDV in different oat samples, nine field-collected oat leaf samples with or without BYDV-like symptoms were analyzed using both RT-RPA and RT-PCR. Both methods generated expected-size amplicons from three of the nine field samples, which indicated that the RT-RPA assay could be used reliably for detecting BYDV infection in field-collected samples (Fig. 4).
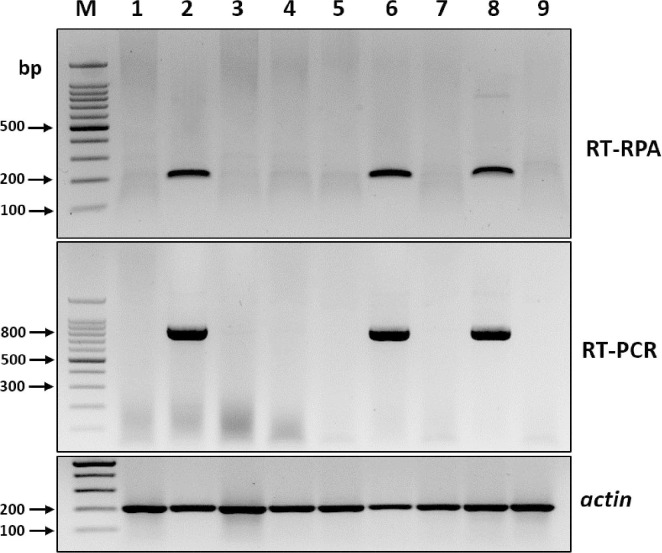
Sample test of barley yellow dwarf virus (BYDV) by reverse transcription recombinase polymerase amplification (RT-RPA). RT-PCR, reverse transcription polymerase chain reaction; M, DNA marker; lanes 1-9, product amplified from nine symptomatic BYDV oat leaves that were collected from the field. An actin was used as an internal control.
The ability of the RT-RPA assay to detect BYDV in R. padi, the main aphid vectors of BYDV, was also investigated. Four aphids collected from BYDV-infected oat leaves were analyzed using both RT-PCR and RT-RPA. The assay used same amount of RNAs. For one aphid sample, the RT-PCR amplified a faint 900-bp band, whereas the RT-RPA assay amplified a clear ~182-bp fragment (Fig. 5). Thus, the RT-RPA assay could be more sensitive than the RT-PCR assay for detecting BYDV in R. padi.

Detection of barley yellow dwarf virus (BYDV) in single aphid by reverse transcription recombinase polymerase amplification (RT-RPA). RT-PCR, reverse transcription polymerase chain reaction; M, DNA marker; lanes 1-4, product amplified from BYDV-harboring (lane 1) and BYDV-free (lanes 2-4) samples. An actin gene from Rhopalosiphum padi was used as an internal control (Wu et al., 2014).
Crop production is expected to increase with increasing global temperatures, but so is the incidence of plant diseases. BYDV, in particular, is considered a serious and widespread threat to cereal grain production, and more importantly, co-infections with other viruses can result in serious cereal disease (Walls III et al., 2019). Therefore, the development of a simple, sensitive, and reliable method for BYDV detection is urgent. In the present study, an RT-RPA assay for the isothermal detection of BYDV in oat leaves and aphids was developed. The assay can be performed rapidly in 5 min and only requires incubation at a single temperature (42°C). The assay was also 100 times more sensitive than RT-PCR, and there was no cross-reactivity with other viruses that are common among cereal crops. The utilities of both the RT-RPA and RT-PCR assays were validated using field samples and aphids. Therefore, RT-RPA could potentially be used for routine field monitoring, which will help farmers to prepare for the spread of the virus (Fig. 6).
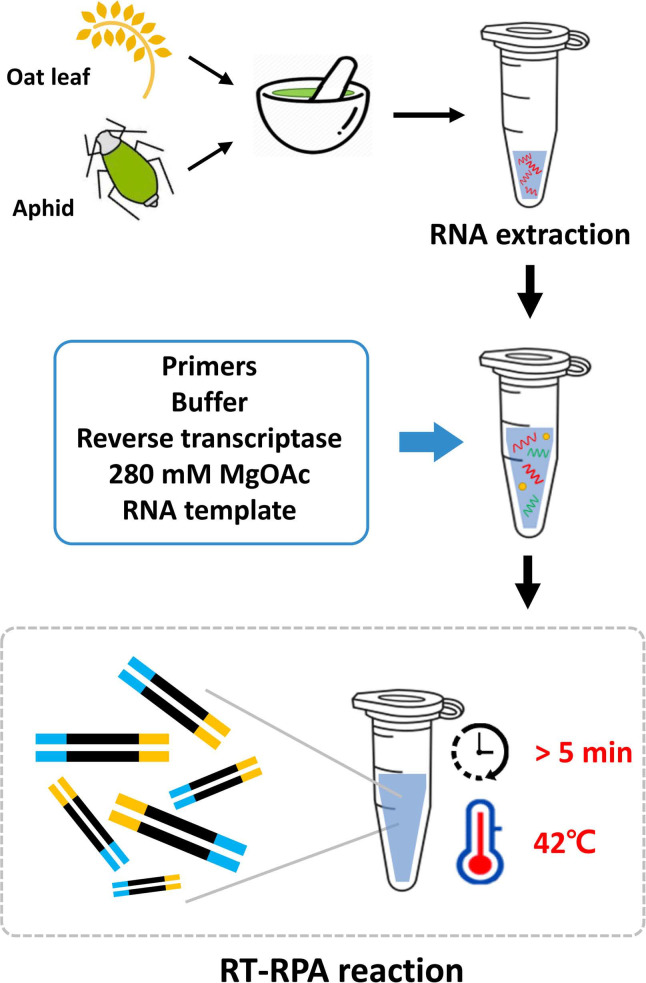
One-step reverse transcription recombinase polymerase amplification (RT-RPA) assay for the detection of barley yellow dwarf virus (BYDV) in oats.
In summary, the present study provides a simple and rapid method for detecting BYDV in oat crops. The RT-RPA assay can be performed rapidly in 5 min, and the resulting product can be visualized using agarose gel electrophoresis. In addition, the assay is over 100 times more sensitive than RT-PCR. Thus, the RT-RPA assay described here represents an effective alternative for detecting BYDV in both oats and aphid vectors, thereby facilitating the restriction on the spread of the virus.
Acknowledgments
This work was carried out with the support of Cooperative Research Program for Agriculture Science and Technology Development (Project No. PJ014995022020) Rural Development Administration, Republic of Korea.

