Update on Distribution and Genetic Variability of Plum pox virus Strains in Bulgaria
Article information
Abstract
Field surveys for Plum pox virus (PPV) infection were conducted in stone fruit orchards all over Bulgaria. In total, 1168 out of 3020 leaf samples from cultivated Prunus spp. and wildly growing P. cerasifera trees reacted positive for PPV in DASI-ELISA with the universal monoclonal antibody (MAb) 5B. Further ELISA analyses showed that 987 and 127 isolates belonged to PPV-M and PPV-D serotypes, respectively. The plum and P. cerasifera showed 82.0% and 50.5% levels of infection, respectively followed by the peach (40.0%) and the apricot (32.0%). Five hundred fifty one PPV isolates were further typed by IC-RT-PCR with PPV-Rec, -M and -D-specific primers, targeting (Cter)NIb-(Nter) CP genome region, as 125 isolates were sequenced. The results revealed the presence of PPV-Rec, PPV-M and PPV-D and mixed infections of these strains. PPV-Rec was the most prevalent strain (49.0%), followed by PPV-M (40.1%), while PPV-D was the less spread strain (8.2%). PPV-Rec was the most common strain in plums, including the eight “old-aged” trees from the region of the first Sharka discovery. PPV-M was the most prevalent strain in peach and apricot. Phylogenetic analyses on (Cter)NIb-(Nter)CP of the isolates were performed. PPV-Rec isolates formed a homogeneous group, while PPV-M isolates split into PPV-Ma and PPV-Mb subgroups. Five separated clades were formed by the analyzed PPV-D isolates. Nucleotide sequences of the partial CP coding region of the analyzed isolates revealed a slightly higher intra-strain genetic variability in PPV-Rec and PPV-M isolates, while that of PPV-D strain isolates was higher from the reported for these strains.
Sharka caused by Plum pox virus (PPV) is one of the most harmful diseases affecting stone (Prunus) fruits crop. Based on the scientific and economic importance PPV is considered as one of the top ten plant viruses (Scholthof et al., 2011). The efficient transmission of PPV by many aphid species in a non-persistent manner, the wide range of isolates differing in their biological, serological and molecular properties and the rare presence of resistance genes within Prunus genes make the implementation of control measures very difficult. Now, Sharka disease is present in Europe, North and Central America, Asia, and North Africa.
To date ten PPV strains as Dideron (D), Marcus (M), Recombinant (Rec), Cherry (C), Cherry Russian (CR), EL Amar (EA), Winona (W), Turkish (T), Ancestral (An) and Cherry Volga (CV) are distinguished (Sheveleva et al., 2018). The above PPV strains differ in their host plant range and pathogenicity on the particular host, antigenic and epidemiological properties and geographical distribution. From all mentioned strains PPV-Rec, PPV-M and PPV-D are widely spread in Europe and Mediterranean basin having an epidemiological importance (Candresse and Cambra, 2006).
Sharka disease described for the first time in Bulgaria (Atanasoff, 1932) has an endemic incidence in the country (Kamenova et al., 2015). From all Prunus species, the plum (Prunus domestica) and sweet cherry (P. avium) are the most cultivated in the country. The peach (P. persica) is also cultivated, mostly in the warmer locations of Southern Bulgaria, while the apricot (P. armeniaca) is the less grown stone fruit species.
In this study, PPV infection rate and strain distribution in the main stone fruit species from all over the country were determined by serological and molecular assays. In addition, wildly-growing P. cerasifera trees were included. Our particular interest was directed also to determine the strain status of “old-aged” PPV infected trees of the local cultivar Kyustendilska sinya in the region of the first detection of Sharka disease.
Materials and Methods
Field surveys
Surveys were carried out in a number of commercial orchards in thirteen different districts of Northern and Southern regions of the country in the period of 2009–2016. Leaf samples were collected from a total of 3020 different stone fruit trees including plum, peach, apricot, sweet cherry, sour cherry and P. cerasifera trees growing along the roads and around the orchards. Five to 10 samples per orchard/tree with or without virus-like symptoms were taken from different parts of the tree crown, placed in an ice bag and kept at 4ºC prior to testing. The positive samples were also dried under CaCl2.
Serological assays
All leaf samples were tested for the presence of PPV by double-antibody sandwich indirect enzyme-linked immunosorbent assay (DASI-ELISA), using the universal monoclonal antibody (MAb) 5B (Agritest, Italy) according to (Cambra et al., 1994). The samples that reacted positive were further analysed with serotype-specific MAbs against PPV-M (Agritest, Italy) and PPV-D strains (Durviz, Spain). Samples with absorbance values (at 405 nm) greater than three times the mean of the absorbance of the healthy controls were considered positive.
Immunocapture reverse transcription polymerase chain reaction (IC-RT-PCR) strain typing
The same extracts tested positive by DASI-ELISA were used for IC-RT-PCR assays. In total 551 isolates (321 from plum, 140 from peach, 78 from apricot and 12 from P. cersifera) were typed with PPV-Rec, -M and -D-specific primers (mD5/mM3, mM5/mM3 and mD5/mD3, respectively) targeting (Cter)NIb-(Nter)CP genome region (Šubr et al., 2004). PCR amplification of PPV isolates with the primers described above gave product as follows: mD5/mM3: 605-bp, mM5/mM3:459-bp and mD5/mD3: 664-bp.
Partial sequencing and phylogenetic analyses
In total 125 isolates (57 defined as PPV-Rec, 46 as PPV-M and 26 as PPV-D) were sequenced in (Cter)NIb-(Nter)CP genomic region. The amplicons obtained were purified (Illusta™, GFX™ PCR DNA Purification kit GF, Healthcare, UK) and sequenced directly in both directions (Microsynth sequencing service, Switzerland or GATC-Biotech, Germany) by priming the reactions with the oligonucleotides used for the PCR. The accuracy of the sequences was validated by BLAST option of the National Center for Biotechnology Information, Bethesda, MD, USA.
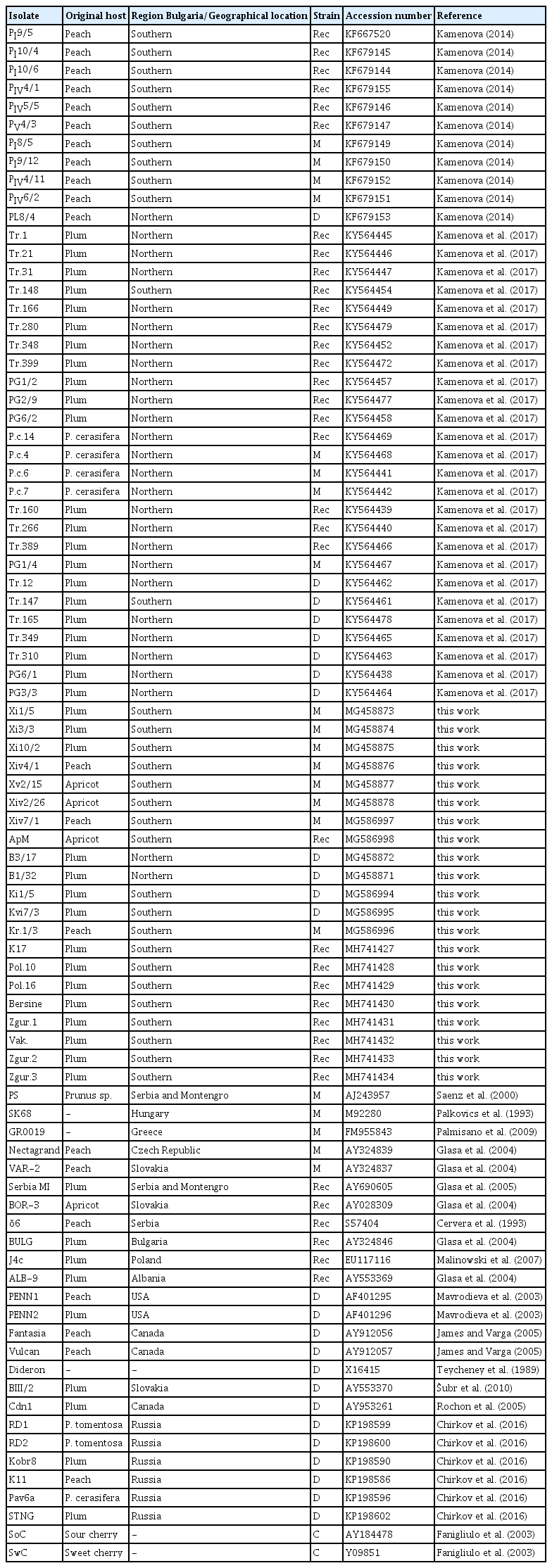
Nucleotide sequences of 67 isolates named as BG (BG01, −02, −03, −04, −05, −06, −07, −08, −09, −10, −11, −12, −13, −14, −15, −16, −18, −19, −20, −21, −22, −23, −24, −26, −27, −28, −29, −30, −31, −32, −33, −34, −35, −36, −37, −38, −39, −40, −41, −42, −43, −44, −45, −46, −47, −48, −49, −50, −51, −52, −53, −54, −55, −56, −57, −58, −59, −60, −61, −62, −63, −64, −65, −66, −67, −68 and −69) are available in the SharCo database (www.sharco.eu). Nucleotide sequences of all other isolates were deposited in GenBank (www.ncbi.nlm.nih.gov) and analyzed together with other PPV CP available gene sequences (Table 1).
Multiple alignments of nucleotide and amino acid sequences of PPV CP gene were carried out using ClustalW (Thompson et al., 1994). The alignments were used to construct phylogenetic trees by the neighbor-joining method, Kimura 2-model of nucleotide substitutions on a computer programme MEGA version 6 (Tamura et al., 2013). Available sequences of 24 isolates corresponding to PPV-Rev, PPV-M and PPV-D (Table 1) from different geographical locations were retrieved from GenBank (www.ncbi.nlm.nih.gov). The divergent C [SoC (AY184478) and SwC (Y09851)] strains were used as out-group.
Results
Strain occurrence and distribution
To represent the different orchard types in the country big and small production orchards and residential gardens with differently aged trees in each of the two regions were visited. From the analyzed 3020 samples from plum, peach, apricot, sweet cherry, sour cherry and P. cerasifera, 1168 (38.7%) were infected with PPV. The highest level of infection was detected in plum (82.0%) and P. cerasifera (50.6%), followed by peach (40.0%) and apricot (32.0%). All samples from sweet and sour cherry tested negative for PPV, nevertheless they were collected from trees growing in close proximity to infected Prunus species.
Of the positive 1,168 samples, 987 and 127 reacted with PPV-M and PPV-D strain-specific antibodies, respectively. Twenty six samples reacted to both PPV-M and PPV-D-specific antibodies. Twenty eight samples that did not react with the above mentioned antibodies, as well as with PPV-C and PPV-EA MAbs were serologically unidentified.
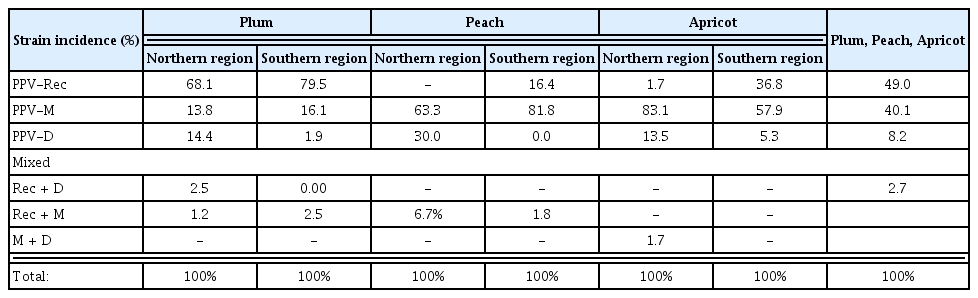
After the initial serological differentiation of M and D isolates, IC-RT-PCR analyses were performed on 551 isolates. The results (Table 2) revealed that PPV-Rec, PPV-M and PPV-D and mixed infections of these strains were present, as the majority of PPV-M serotyped isolates were in fact PPV-Rec (65%) (PPV-Rec isolates are serologically related to PPV-M). PPV-Rec was the most prevalent strain on plum found in 79.5% and 68.1% of the analyzed samples from Southern and Northern regions, respectively. Recombinant isolates were present in most plum samples collected from eleven different districts in both regions, with exception of two (one in each of the two regions). PPV-M was distributed in almost equally extent in both regions, while PPV-D predominated in the Northern region.
Unlike the plum, the peach and apricot were infected mainly by PPV-M strain with infection rates of 81.8% and 63.3% (peach) and 57.9% and 83.1% (apricot) in the Southern and Northern regions, respectively. PPV-Rec in peach found in two commercial orchards in a location close to the Greek border (Southwest) was the first detection of natural peach infection by this strain (Kamenova et al., 2011).
PPV-D was the less spread strain in all tested Prunus species in Bulgaria, accounting for 8.2% of all detected infections. Concerning its presence in the different Prunus species it was found in peach in only one orchard in Northern region, where nine trees out of eleven tested were positive for this strain and in single trees in several plum orchards (totally 6 in the two regions).
The rate of mixed infections (Rec + D and Rec + M) compared to the single infections with each of the three strains was relatively low (total 6.2%, 8.5 in plum and peach, respectively). Mixed infection of M + D at a rate of 1.7% was found only in apricot infected trees.
As a total, PPV-Rec was the most prevalent strain (49.0%) in Bulgaria overall, followed by PPV-M (40.1%), while PPV-D was the less spread strain (8.2%) in tested Prunus species. A clear prevalence of PPV-Rec in plum and PPV-M in peach and apricot under natural condition was observed.
Establishment of PPV-Rec as the most spread strain on plum motivated further investigation. The intent was to determine the strain status of PPV infected “old-aged” trees of the local variety Kyustendilska sinya from the region of first Sharka detection. In total eight trees from three locations of the region were analyzed. The sampled trees were more than 70 years old (according to the owners), as in one of the locations the trees were isolated and completely abandoned, swallowed by the growing in the terrain forest species. To prevent the risk of inaccurately strain characterization by single leaf sampling, four samples were collected from the four directions (east, west, south and north) of the canopy of each tree. These samples were serologically and molecularly typed. All 32 samples reacted with only PPV-M-specific antibodies. In seven of the analyzed trees (28 samples) only PPV-Rec was detected, and in one tree (4 samples) mix of Rec + M was found.
Genetic variability of detected PPV strains
PPV-Rec
The nucleotide sequences of 57 isolates covering the (Cter)NIb-(Nter)CP coding region were aligned with six GenBank-deposited European Rec isolates as BOR-3, J4c, ŏ6, Serbia-MI, ALB-9 and BULG using ClustalW (Thompson et al., 1994). Generated phylogenetic tree (Fig. 1) confirmed the results of the RT-PCR typing and showed that the analyzed isolates formed a monophyletic group. The isolates from peach generated a separate clade, while the isolates from plum, apricot and P. cerasifera dispersed in common ones. The isolates from “old-aged” trees separated in different clusters, as four of them (Zgur.1, Zgur.2, Zgur.3 and K17 - all from one location of Kyustendil region) were closely related with two isolates from apricot and plum (ApM and BG14, respectively) from different locations. Two others “old-aged” isolates (Bersine and Pol.16) took positions among many isolates from plum, while isolate Vak was in a cluster with several reference isolates (ŏ6, Serbia-MI and Jc4) with origin from Serbia and Poland. One of the analyzed plum isolates (BG45) clustered with two reference isolates, BOR-3 and ALB-9 from apricot and plum, respectively.

Phylogenetic tree of Plum pox virus strain Rec isolates from Bulgaria reconstructed from (Cter)NIb-(Nter)CP genomic region. The scale represents a distance of 0.05 substitutions per site. The tree was reconstructed by neighbour joining using the Kamura-2 parameter model of evolution. Only bootstrap values > 70% are shown. The isolates sequenced in this study are boldface. After the name of the isolate the original host is shown as plplum, pe-peach, ap-apricot and p.c-P. cerasifera.
The number of variable nucleotides among studied isolates was counted directly after alignment and obtaining consensus. This number ranged from 0 (among several peach isolates, among several isolates from “old-aged” trees and between these isolates and several others plum isolates) to 12 (between isolates from peach and plum). The average nucleotide genetic diversity among isolates from each host species and between isolates from the different species is shown in Table 3. The largest intra-strain diversity was observed among the reference isolates (0.022 ± 0.005) originating from several countries as Serbia (ŏ6 and Serbia-MI), J4c (Poland), Slovakia (BOR-3) and Albania (ALB9), while the lowest one among the eight “old-aged” isolates (0.010 ± 0.004 – not shown) and the isolates from peach (0.012 ± 0.003). The mean nucleotide divergence for the isolates from the different Prunus species was the highest between peach and reference and peach and plum isolates (0.033 ± 0.007, 0.029 ± 0.006), respectively.
The deduced (Cter)NIb-(Nter)CP amino acid sequences of PPV-Rec isolates were compared (not shown). The results obtained showed identity levels among plum isolates of 93–100% (from 0 to 7 different amino acids). The isolates from peach were 96–100% identical (from 0 to 4 differences), while the identity between plum and peach isolates was 92–98%. This identity level was lower than the reported of 95–100% (Matic et al., 2011) for the compered 19 PPV-Rec isolates available in GenBank most probably due to the absence of peach isolates in the comparison. The level of amino acid identity among the “old isolates” was 97–100%, as two of these isolates (Bersine and Pol.16) were 100% identical to a number of isolates as BG21, −24, −25, −27, −33, −36, −39, −47, −49 and Tr.280, and to the reference isolate BOR-3.
The amino acid comparison showed also that the coat proteins of studied isolates started with the amino acid alanine (A) which is valid for PPV, with exception of several isolates (Kamenova et al., 2001; Maiss et al., 1995) and that all of them contained 11DAG13 motif, considered as essential for aphid transmission. In all analyzed isolates the “typical” sequence of the universal epitope 94DRD-VDAG100 was present (Candresse et al., 2011) and they were recognized by the MAb5B.
PPV isolates differ in the mobility of their capsid protein (CP) in SDS-PAGE (Adamolle, 1993; Bousalem et al., 1994), as PPV-Rec isolates have shown to migrate in a double-band form (Šubr and Glasa, 1999; Šubr et al., 2007). Amino acid comparison of our PPV-Rec isolates showed that several plum isolates (BG23, Tr.21, Tr.31, Tr.148, Tr.399, PG1/2 and PG6/2) showed the presence of arginine (R) (66 position in SPISR motif), instead of glycine (G) (SPISG) in N-terminal region, thus suggesting the formation of a single-band in SDS-PAGE analyses as reported for isolate BULG (Šubr et al., 2010). Using site-directed mutagenesis the authors have demonstrated that glycine (66G) in the CP has resulted in the double-band phenotype, while arginine (66R) has led to a single-band CP pattern.
PPV-M
The obtained nucleotide sequences of 46 isolates were aligned with the reference isolates SK68 and PS, as representatives of two reported Ma and Mb clades (Dallot et al., 2011) and three more isolates (VAR-2, Nactagrand and GR0019) from different geographical locations, as the first two originated from Eastern European countries (Czech Republic and Slovakia, respectively), while GR0019 was from Greece. The results of phylogenetic analyses (Fig. 2) demonstrated that most of the isolates (totally 36) belonged to PPV-Mb clade, where they formed separate clusters and few more isolates (totally 8) were in Ma clade. One of studied peach isolates, named Kr.1/3 from an orchard located very close to Turkish border, stand out separately from Ma and Mb clades. This isolate showed 91%–96% identity with all M isolates (from 3 to 19 nucleotide differences).

Phylogenetic tree of Plum pox virus strain M isolates from Bulgaria reconstructed from (Cter)NIb-(Nter) CP genomic region. The scale represents a distance of 0.05 substitutions per site. The tree was reconstructed by neighbour joining using the Kamura-2 parameter model of evolution. Only bootstrap values > 70% are shown. The isolates sequenced in this study are boldface. After the name of the isolate the original host is shown as pl-plum, pe-peach, ap-apricot and p.c-P. cerasifera.
In general no host-based and/or the local region (Northern, Southern) clustering within each clade was observed. For example, part of the isolates from peach collected from orchards close to the Greek border (Petrich and Xaskovo districts) belonged to PPV-Mb.
The mean nucleotide divergence among PPV-Mb clade isolates was lower (0.019 ± 0.005), compared to that of PPV-Ma clade isolates (0.030 ± 0.005). The mean nucleotide divergence between Ma and Mb clade isolates was 0.027 ± 0.005. In general the mean nucleotide divergence among all Bulgarian PPV-M isolates was higher (0.020 ± 0.004), than the reported for PPV-M isolates (0.013 ± 0.002) in CP coding region (Dallot et al., 2011).
The deduced amino-acid sequences of all Bulgarian M isolates were 91–100% identical, as the lowest identity (91%) was found between isolates belonging to Ma and Mb group [(BG64Ma X3/6Mb – difference of 11amino acids) and (BG64Ma and BG07Mb - 12 different amino-acids). In fact, part of the isolates (BG10, BG37 and BG64) belonging to Ma clade showed arginine (R) at position 4 (Nter CP) present in PS isolate (Mb), instead of glutamic (E) present in SK68 isolate, while 14 isolates belonging to Mb clade, as well as isolate Kr.1/3 showed at the same position the presence of lysine (K2814) reported by (Glasa et al. 2002) as conserved only within recombinant sequences and found also by Deborré et al. (1995) in two more PPV-M isolates.
Isolate Kr.3/1 was 92–97% identical to the analyzed isolates. BLAST analyses of this isolate with PPV-M isolates from GenBank showed its highest identity (98% and 99%) with shch and cr5 isolates, both from Greece (CUU47629.1 and CUU47620.1), respectively.
PPV-D
An analysis based on alignments of the nucleotide sequences of the (Cter)NIb-(Nter)CP genomic region of 26 isolates (Fig. 3) showed that they formed five separate clusters and did no group with any of the reference isolates. Four of the analyzed isolates (two, respectively from apricot and plum) all from the extreme eastern locations of the Northern region dispersed between reference isolates with origin from Russia (Kobr8, RD1, RD2).
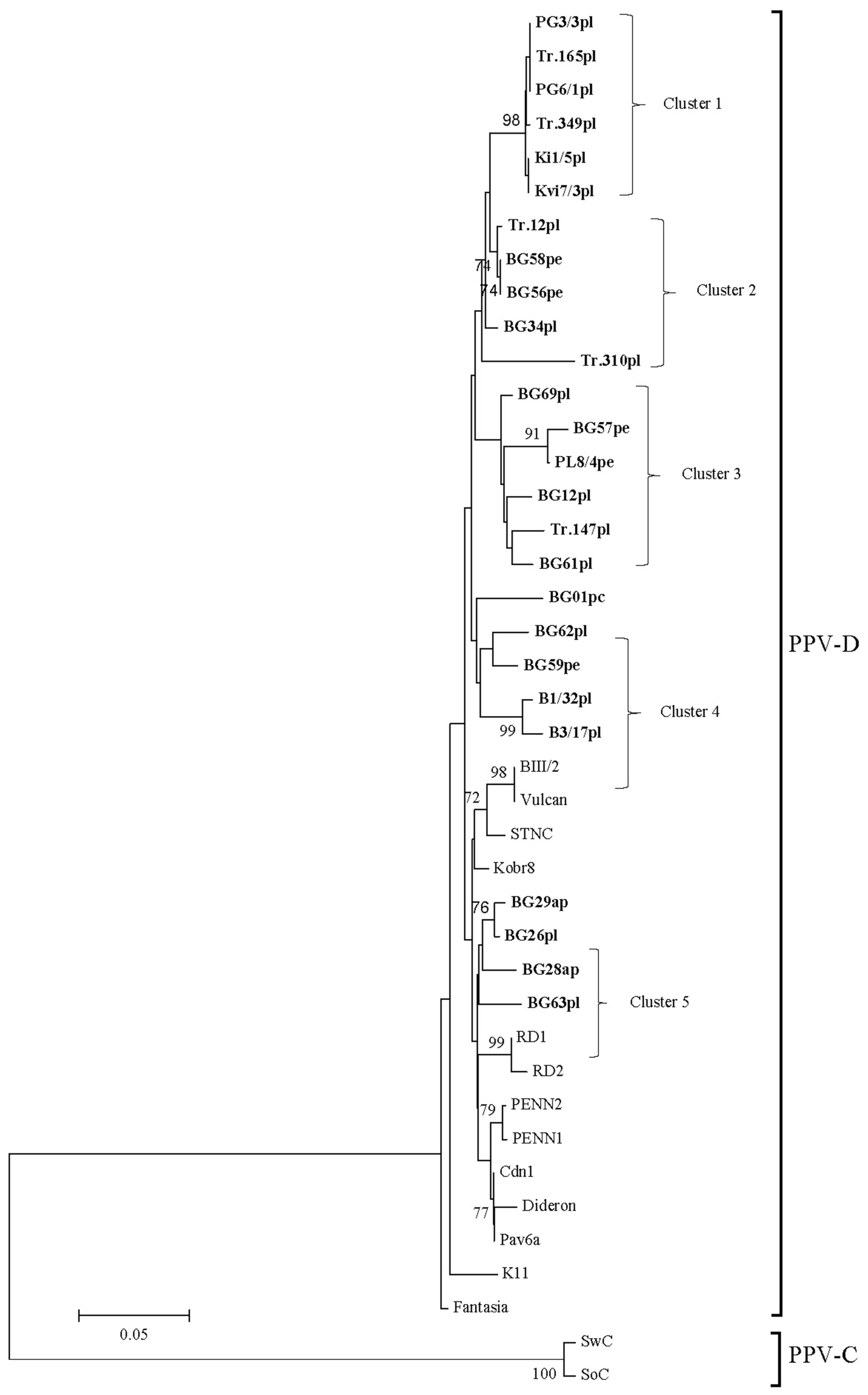
Phylogenetic tree of Plum pox virus strain D isolates from Bulgaria reconstructed from (Cter)NIb-(Nter)CP genomic region. The scale represents a distance of 0.05 substitutions per site. The tree was reconstructed by neighbour joining using the Kamura-2 parameter model of evolution. Only bootstrap values > 70% are shown. The isolates sequenced in this study are boldface. After the name of the isolate the original host is shown as pl-plum, pe-peach, apapricot and p.c-P. cerasifera.
The mean intra-strain nucleotide divergence level among Bulgarian PPV-D isolates was much higher (0.042 ± 0.005), than the reported (0.013 ± 0.001) for the full CP genomic region (Dallot et al., 2011).
A comparison of the deduced amino-acid sequence residues in the CP region of the analyzed isolated revealed identity of 89–100%, as two of the isolates B1/32 and B3/17 from plum showed the lowest one (89–92%) to all other isolates (from 8 to 12 different amino-acids). While 96–99% amino acid identity of PPV-D strain isolates was reported by Matic et al. (2011), blast analyses of B1/32 and B3/17 isolates with available in GenBank PPV-D isolates showed identity of only 88% with Pakistan-D isolate (ABD85470.1) and isolates from Chile and USA (AAL35270.1; AAL35274.1; AEY82641.1 ).
During the initial serological differentiation performed with monoclonal antibodies AL and 4DG5 specific for the PPV-M and PPV-D serotypes, respectively no reaction was received with two plum isolates named B1/32 and B3/17 (Northern region), nevertheless their positive reactions with the universal 5BMAb (absorbance values at 405 nm −0.193 and 0.546, respectively). The 664-bp products were obtained by the followed RT-PCR analyses using PPV-D-specific primers mD5/mD3. Comparison of amino acid sequences of these two isolates with that of other Bulgarian and reference PPV-D isolates revealed the presence of the “typical” sequence of the universal epitope 94DRDVDAG100 (Candresse et al., 2011), responsible for their recognition by 5B. However, while the compared PPV-D isolates contained the putative 4DG5-binding sequence 49FTPAT-TQPATKPVS62, B1/32 and B3/17 showed one common aa substitutions as V61A. Additionally, isolate B1/32 and B3/17 contained one more substitution, including T58I and K60E, respectively. It should be noted that the mutation K60E in B3/17 isolate has been shown to abolish the reactivity of the Bosnian isolate BOS49PL (AJ49999) (Matic et al., 2006). Most probably the mutation T58I in isolate B1/32 could be responsible for the lack of recognition by 4DG5, as in the case of the Crimean K27 (KR028387) and the Canadian Vulcan (AY912057) isolates reported to shown T58P and T58A, respectively (Chirkov et al., 2016). The results obtained with B1/32 and B3/17 provided evidence that the amino acid substitution at positions pointed above affected their recognition by antibody 4DG5 in agreement with the reported by Candresse et al. (2011) and Chirkov et al. (2016).
Discussion
PPV strain presence in cultivated Prunus species all over the country and their genetic variability were studied. Serological and molecular typing of large number of PPV isolates from plum, peach, apricot and P. cerasifera confirmed the spread of PPV-Rec, PPV-M and PPV-D strain (Kamenova et al., 2015).
Sweet cherry being the most commonly grown stone fruit species in the country was found not infected by PPV, consistent with its previously reported absence in sweet and sour cherry in the country (Kamenova et al., 2013). According to Sihelska et al. (2016) PPV-C and PPV-CR strains have the strictest linkages with sweet and sour cherry under natural condition. With the experimental transmission of PPV-Rec and PPV-M strains to mahaleb cherry (Milusheva, 2008), and reported PPV-M infection of cherry in Croatia (Mikec et al., 2008) and in the Czech Republic (Navrátil et al., 2008), the possibility of adaptation of other PPV strains to these Prunus species exists.
As the second most grown stone fruit species the plum had the highest infection rate (82.0%), followed by the lesser grown peach and apricot.
While the strain status of PPV on cultivated species in Bulgaria was more or less determined in this study, this of the ornamental and wild-growing Prunus species is still scarce and insufficient. It should be noted that most of the samples that were not serologically determined originated from P. cerasifera. Further molecular strain typing and sequence analyses of PPV isolates from P. cerasifera are ongoing.
Variability of PPV isolates can be studied by a number of advanced analytical methods, including identification by DASI-ELISA with monoclonal antibodies to D, M, C and EA strains, molecular typing by RT-PCR with strain-specific primers and sequence analyses of CP gene, followed by phylogenetic analyses (Šubr and Glasa, 2008).
The results from DASI-ELISA test proved the reliability of the universal MAb 5B in detection of PPV strains (Candresse et al., 2011). From the found 1168 PPV positive samples, twenty eight (2.4%) escaped the serological strain differentiation with the use of PPV-M, -D -C and -EA MAbs. The affinity of MAbs may be influenced even by a singly amino acid change, as in the recent reported D96E mutation in the 5B universal epitope in a range PPV isolates from sour cherry in Russia (Sheveleva et al., 2018). This statement was confirmed in the case with two of the isolates (B3/17 and B1/32) not recognized by the use of PPV-D strain specific Mab, but defined to belonged to this strain by the followed molecular and sequence analyses. Thus the potential escape of some PPV isolates from typing with MAbs emphasizes the importance of molecular strain typing (Candresse and Cambra, 2006).
Performed RT-PCR analyses with strain-specific primers targeting (Nter)NIb-(Cter)CP genomic region allowed PPV-M and PPV-Res strain isolates, which are serologically related to be distinguished. PPV-Rec, PPV-M and PPV-D strain were detected as already reported (Kamenova et al., 2015), thus confirming that these three strains are the major ones from a global point of view (Šubr and Glasa, 2013). A different relative ratio of the strains was found as PPV-Rec and PPV-D were the most and less spread strains, respectively (6Rec to 1D; 1.2Rec to 1M and 4.8M to 1D). While both PPV-Rec and PPV-M were found almost evenly spread in the two regions, PPV-D was registered in a higher extent in the Northern region.
Given the wide distribution and prevalence of PPV-Rec strain in plum, which is one of the most cultivated Prunus species in the country, this strain could be accepted as the major one in Bulgaria. In general PPV-Rec is spread in central and south-eastern Europe, PPV-M is found mainly in southern and central Europe and Turkey, while PPV-D is the strain with the globally broadest spread, as it has been found in Europe, both Americas and several Asian states (Sihelska et al., 2016). The finding of our study showed also a clear prevalence of PPV-Rec in plum and of PPV-M in peach, in correlation with the reported host preference of these isolates (Sihelska et al., 2016). PPV-Rec isolates were reported exclusively spread in plums, while PPV-M isolates were generally linked to the severe epidemics of peach (Šubr and Glasa, 2008). In our study PPV-D was found in only one peach orchard, and in single trees in several plum and apricot orchards. Despite the small number of PPV-D isolates found in Bulgaria, we assume that the plum and apricot are also the preferable hosts for this strain under the ago-climatic conditions in Bulgaria, as reported by Sihelska et al. (2016). The current widespread distribution of PPV-Rec isolates in plums in Bulgaria is a proof both for their good adaptation to this species, and for the existence of rapid and effective mechanisms in their short- and long-distance distribution (found in almost all districts). This situation in Bulgaria was most probably favored by the absence of strict eradication of infected trees, especially in the first years after discovery of the virus nature of Sharka disease. Tracking the spread of PPV-Rec and PPV-D strains in an experimental plum orchard in Bulgaria, PPV-Rec was determined as competitive and endemic strain (Kamenova et al., 2017), similarly to the reported from Serbia (Jevremović, 2012).
No difference on PPV-Rec presence in plums depending on the age of the trees/orchards was found. PPV-Rec was the most prevalent strain on this species overall in the country, and was found in all eight “old-aged” plum trees in the region of the first PPV detection. These results showed a strong link between PPV-Rec and plums and proved unambiguously its long-standing presence in the country, overlooked for a long period of time due to the limitation of serological tests to discriminate M and Rec isolates. It should be noted that in comparison with PPV plum infections, such in peach trees in Bulgaria was found relatively later (Yankulova et al., 1990), nevertheless the numerous surveys carried out by the authors in the main peach-growing regions. An explanation for the observed almost 50 year’s escape of peach infection in the country could be connected with present finding for the domination of PPV-Rec isolates. While widely spread on plums, PPV-Rec was less frequently associated with peach under natural conditions (Garcia et al., 2013). Reported presence of PPV-Rec in several peach orchards in southwest Bulgaria, however, could be some indication for its possible larger adaption to this species under the specific agro-climatic conditions of the region (Kamenova, 2014).
Sequence comparison and construction of the phylogenetic trees provide the best depiction of the relationships among isolates (Candresse and Cambra, 2006). Phylogenetically the studied Bulgarian PPV-Rec isolates were closely related among them and to the reference isolates, while PPV-M isolates split into two subgroups, PPV-Ma and PPV-Mb found to be associated with their geographical origin. It is accepted that PPV-M isolates from eastern European countries as the Czech Republic, Serbia, Slovakia and Bulgaria belong mainly to the PPV-Mb clade, while the isolates from countries as France, Italy Cyprus and Greece belong to the PPV-Ma clade (Dallot et al., 2011). Our results showed the presence of both PPV-Ma and PPV-Mb isolates in Bulgaria, with clear prevalence of isolates belonging to Mb group. Mb isolates were found in the two regions in infected Prunus spp., as all peach isolates (excluding isolate BG10) from Petrich district which is very close (few kilometers) to the Greek border belonged also to Mb group. A possible explanation for the prevalence of PPV-Mb isolates in Bulgaria could be looking in the historical trade exchanges between Bulgaria and Eastern European countries in the years before 1990. Phylogenetically studied PPV-D isolates showed 99% bootstrap support, without geographical location/or host-associated clustering. It should be pointed, that the number of studied PPV-D isolates was relatively low, and in addition almost all isolates originated from only several districts in Northern region.
Although genetic variability of PPV isolates from many European countries has been well studied, this of Bulgarian isolates was poorly investigated. Nucleotide sequences of the partial CP coding region of the analyzed 125 isolates revealed a slightly higher intra-strain genetic variability in PPV-Rec (0.018 ± 0.003) and PPV-M (0.020 ± 0.004) isolates from the reported for these strains (0.015 ± 0.002 and 0,013 ± 0.002, respectively) (Dallot et al., 2011), while PPV-D strain isolates exhibited much higher intra-strain variability (0.042 ± 0.005) in comparison to the reported (0.024 ± 0.002; 0.013 ± 0.001) (Dallot et al., 2011; Glasa et al., 2004). In general agronomical and biological factors can facilitate the emergence of genetic diversity in PPV, as scion-rootstock combination, movement of infected plants, introduction of new species and last but not least vector-associated selection (James et al., 2013). According to Garcia et al. (2013) the observed by Jridi et al. (2006) and Predajňa et al. (2012) substantial intra-isolate variability within single Prunus tree is an indication for the dynamic structure and heterogeneous nature of PPV populations.
Calculated intra-strain nucleotide diversity of the “old-aged” isolates (0.010 ± 0.004) was the lowest one and not consistent with the reported of 0.022 ± 0.003 for several isolates from former Yugoslavia (Glasa et al., 2005). Based on the statements of Garcia-Arenal et al. (2001) and Wei et al. (2009), that the older population are expected to accumulate more genetic diversity than the younger ones, Glasa et al. (2005) suggested that the former Yugoslavia is the original center of dispersion of PPV-Rec isolates. Indeed the first reported PPV recombinant isolate ŏ6 originated from Serbia (Cervera et al., 1993) and on that base the Balkans were suggested as the center of the origin of PPV-Rec isolates (Glasa et al., 2005). The observed low diversity among studied “old-aged” Bulgarian isolates, however, could due to some other effects, as different cultural practices, different host range or genetic bottlenecks during aphid transmission of the virus population as pointed by (Ali et al., 2006; Glasa et al., 2007). According to Wei et al. (2009) the history of a disease may, but may also not be consistent with the degree of genetic variability and thus to be relevant to the history and origin of PPV-Rec isolates.
The finding of PPV-Rec isolates considered to be recombinant between D and M isolates (Glasa et al., 2004) in all eight sampled “old-aged” trees and in mixed infection with PPV-M in one of the tree of the local variety Kyustendilska sinya in the area if the first Sharka discovery aroused some questions. Could be PPV-Rec an autochthonous strain in Bulgaria? Could PPV-Rec isolates be an ancestral group, while either PPV-M or PPV-D to have resulted from it? In support to this though indirect is the fact that PPV-Rec predominated in all regions in the most infected Prunus species, while the level of mixed infections of the three strains found in the country, particularly in plum infected trees (3.1%) and totally in all infected Prunus species was low (2.4%), thus reducing the possibility PPV-Rec to has derived from a recombination between PPV-M and PPV-D. By the results obtained it is not possible to answer to the above questions. However, the feasibility of such an assumption should not be excluded. Experiments with inter-strain mixed infections both in woody and herbaceous hosts, have had not leaded to the recombination de novo (Capote et al., 2006). According to Šubr et al. (2008) the expansion of PPV-Rec is rather consequence of its high competitiveness than a “hot-spot” recombination in the PPV genome. It is clear that analyses of more “old-aged” isolates, encompassing the full-length genome and possibly from more countries is needed.
In this study, the current state of knowledge about PPV strain presence and their distribution on the main Prunus species all over Bulgaria was broadens with a range of serological and molecular analyses. PPV-Rec, PPV M and PPV-D isolates were found in cultivated Prunus species. A presence of PPV-Rec isolates in “old-aged” trees in the region of the first Sharka discovery, their major prevalence in plums and relatively low genetic diversity was shown. The finding of both PPV-Ma and PPV-Mb isolates is a reflection of the enlarged trade of infected planting material with Eastern and Western-European countries. The substantial diversity of Bulgarian PPV-D isolates could be result from the specific host species/cultivars grown in the country, the impact of the vectors and/or of agro-climatic conditions. In general the results presented here contribute to the expanding of the knowledge of PPV genetic variability and characterization, thus affording to better understand the epidemiology of the pathogen which is among the first steps in implementation of effective Sharka - control strategies.
Acknowledgments
Part of the research leading to these results was founded by the EU Seventh Framework Programme (FP7/2008-3013) Grant Agreement No. 204429 SharCo project.