Occurrence of Squash yellow mild mottle virus and Pepper golden mosaic virus in Potential New Hosts in Costa Rica
Article information
Abstract
Leaf samples of Solanum lycopersicum, Capsicum annuum, Cucurbita moschata, Cucurbita pepo, Sechium edule and Erythrina spp. were collected. All samples were positive for begomoviruses using polymerase chain reaction and degenerate primers. A sequence of ∼1,100 bp was obtained from the genomic component DNA-A of 14 samples. In addition, one sequence of ∼580 bp corresponding to the coat protein (AV1) was obtained from a chayote (S. edule) leaf sample. The presence of Squash yellow mild mottle virus (SYMMoV) and Pepper golden mosaic virus (PepGMV) were confirmed. The host range reported for SYMMoV includes species of the Cucurbitaceae, Caricaceae and Fabaceae families. This report extends the host range of SYMMoV to include the Solanaceae family, and extends the host range of PepGMV to include C. moschata, C. pepo and the Fabaceae Erythrina spp. This is the first report of a begomovirus (PepGMV) infecting chayote in the Western Hemisphere.
Latin America and the Caribbean Islands posses a great diversity and incidence of begomoviruses (Morales, 2010). Since 1950, weeds of the Euphorbiaceae (Costa and Bennett, 1950) and Malvaceae (Costa and Carvalho, 1960) families have been identified as primary sources of inoculum for the transfer of these viruses to nearby crops. However, it was not until the 1970s that the first important losses in production due to viral infections caused by this group of pathogens were reported in crops such as cotton (Gossypium hirsutum and G. barbadense) and bean (Phaseolus vulgaris) (Morales, 2010).
The genus Begomovirus (Geminiviridae) is characterized by a circular single-stranded DNA genome, encapsidated in twinned icosahedral particles. Begomoviruses of New World have bipartite genomes. The DNA-A component include several open reading frames (ORFs) related to replication, encapsidation, movement and transcriptional activation of virion sense genes (Harrison, 1985). The DNA-B component has two ORFs implicated in viral systemic movement (Etessami et al., 1988).
The begomoviruses are recognized as infectious agents causing economic losses in crops in tropical and subtropical zones worldwide (Dhaliwal et al., 2010; Navas-Castillo et al., 2011; Rojas et al., 2005). An important group of begomoviruses in America are those infecting cucurbits. Among the most prominent are Squash leaf curl virus (SLCV), Squash mild leaf curl virus (SMLCV), Cucurbit leaf crumple virus (CuLCrV), Squash yellow mild mottle virus (SYMMoV) and Melon chlorotic leaf curl virus (MCLCuV) which cause economic losses in Cucurbita moschata, C. pepo, C. maxima, Cucumis melo and Citrullus lanatus (Idris et al., 2008; Karkashian et al., 2002; Morales, 2010). Since these viruses can infect the same plant host species, mixed infections have been found in which rearrangements are formed by the encapsidation of DNA-A of one virus species with DNA-B of a different virus species. The degree of virulence depends on the species that are mixed and on the host plant (Idris et al., 2008).
In the Western Hemisphere, the first reports of begomoviruses in pepper (Capsicum spp.) crops were in the United States (Brown et al., 1989; Brown and Poulos, 1990) and Mexico (Leal and Quintero, 1989). Currently, Pepper golden mosaic virus (PepGMV) is one of the most important species (Brown et al., 2005) affecting Capsicum annuum (bell pepper), with a host range that also includes Solanum lycopersicum (tomato) (Nakhla et al., 1994; 2005; Rojas et al., 2000; 2005) and some cucurbits (Ala-Poikela et al., 2005). Other viruses detected in pepper as well as tomato include Pepper huasteco yellow vein virus (PHYVV) (Morales et al., 2005), Tomato leaf curl Sinaloa virus (ToLCSinV) (Brown et al., 1993), Tomato severe rugose virus (ToSRV) (Bezerra-Agasie et al., 2006; Nozaki et al., 2006), Tomato yellow leaf curl virus (TYLCV) (Ascencio-Ibáñez et al., 1999; Quiñones et al., 2001) and Potato yellow mosaic virus (PYMV) (Umaharan et al., 1998).
Begomoviruses identified in Costa Rica to date include PepGMV in pepper (Lotrakul et al., 2000) and tomato (Nakhla et al., 2005), ToLCSinV (Idris et al., 1999; Nakhla et al., 2005) and Tomato yellow mottle virus (ToYMoV) (Hilje et al., 1993; Nakhla et al., 1994) in tomato. Squash yellow mild mottle virus (SYMMoV) has been detected in Cucurbita moschata (squash), Citrullus lanatus (watermelon), Carica papaya (papaya) (Karkashian et al., 2002) and bean (Karkashian et al., 2011). There are also reports of PepGMV (Hilje et al., 1993), Bean golden mosaic virus (BGMV) (Gámez, 1971) and Calopogonium golden mosaic virus (CaGMV) (Díaz et al., 2002) in bean.
The Cartago province, one of the most important areas of vegetable production in Costa Rica, has been studied continuously. Research in this region has shown the possibility of an amplified host range for these viruses and the potential emergence of new virus species from the recombination of existing species.
The objectives of this research were to identify and determine the sequence variability of begomoviruses in Cartago in tomato, pepper, squash, Cucurbita pepo (zucchini, scaloppini), Sechium edule (chayote) and Erythrina spp., and to determine phylogenetic relationships between these and previously described sequences.
Materials and Methods
Sample collection and total nucleic acid extraction to obtain begomovirus DNA.
A total of 15 leaf samples from tomato, pepper, cucurbits and Erythrina spp. were collected in the Cartago province in 2007 and 2008. Samples were stored at −70 °C. The protocol described by Dellaporta et al. (1983) was followed to obtain total nucleic acid.
Rolling circle amplification (RCA) and polymerase chain reaction (PCR) for detection of begomoviruses.
Total DNA purified from 15 leaf samples were amplified by RCA (Fire and Xu, 1995) using the IlustraTM TempliPhi Amplification Kit (GE Healthcare, UK). RCA products were reamplified by PCR using degenerate primers AV494 and AC1048 (Wyatt and Brown, 1996) for the core of capsid protein gene (AV1) to confirm the presence of begomovirus infection. The final volume of the reactions was 25 μl containing: 1× reaction buffer, 2.5 mM MgCl2, 250 μM dNTPs, 0.1 μM each primer, 5U/μl Dream Taq DNA polymerase (Fermentas, Hanover, MD) and 2 μl viral DNA amplified by RCA. The PCR program used the conditions defined by Rojas et al. (1993).
PCR for molecular characterization of begomoviruses.
RCA products of 14 samples were reamplified by PCR using degenerate primers PAC1C1960 and PAV1v722 (Zhou et al., 1997), which correspond to a region of ∼1300 bp of the DNA-A. This region will be referred herin as DNA-A bottom and it includes complete nucleotide sequences of the replication enhancer protein (AC3) and transcriptional activator protein (AC2). Additionally, the amplicon also contains a partial sequence of the replication associated protein (AC1) and coat protein (AV1) genes. PCR reactions were carried out as described previously by Rojas et al. (1993).
Electrophoretic analysis.
RCA and PCR products were analyzed by electrophoresis in TopVision 1% agarose gels (Fermentas) in 1× TAE buffer at 100 V for 40 min. The O’GeneRulerTM 100 bp Plus DNA Ladder and the O’RangeRuler 500 bp DNA Ladder (Fermentas) were used as molecular weight markers. The amplicons were stained with ethidium bromide (1 mg/ml) in water, visualized with ultraviolet light (TFX-20.M, Vilber Lourmat), and photographed with a digital camera (Kodak EDAS 290).
Sequencing and phylogenetic analysis.
The DNA-A bottom from the 14 previously amplified isolates and the AV1 gene from the Cha157 was sequenced. Sequencing was carried out with a BigDye Terminator 3.1v (Applied Biosystems, Foster City, CA) and the automated sequencer Genetic Analyzer 3130, AB (Applied Biosystems, USA). The BLASTn algorithm (Altschul et al., 1990) was used to compare nucleotide sequences of the amplified PCR products with sequences available in the GenBank database. The sequences of the 15 samples begomovirus-positive samples obtained in this study and those from GenBank that shared the greatest nucleotide identity with our samples,were aligned using the Clustal W algorithm (Thompson et al., 1994). Phylogenetic trees were constructed with the MEGA program version 5 (Kumar et al., 2008), using the nucleotide distances obtained and neighbor-joining phylogenetic analysis (Kimura, 1980). Internal branch robustness was estimated by bootstrap analysis with 1000 replicates.
Results
The symptoms observed in plants infected with begomoviruses in this study included yellow mosaic, yellow veins, changes in leaf size and morphology, leaf thickening, thickened vein, and yellow wrinkled leaf margins (Fig. 1). Symptomatic leaf samples detected with begomovirus were collected from tomatoes, peppers, cucurbits and Erythrina spp. In addition, begomovirus infection was detected in one asymptomatic zucchini sample.

Symptoms in (A) Sechium edule (chayote), (B) Erythrina spp. (poro), (C) Solanum lycopersicum (tomato), (D) Capsicum annuum (bell pepper), (E) Cucurbita moschata (squash) and (F) Cucurbita pepo (scallopini), infected with whitefly-transmitted viruses.
RCA products of 14 samples reamplificated by PCR using primers targeted to the DNA-A bottom of the viral genome yielded products circa 1,300 bp for all samples (Fig. 2). These 14 amplicons and the amplicon corresponding to the coat protein (AV1 gene) from the chayote leaf sample (Cha157) were sequenced. Sequences of ∼1,100 bp were obtained from the genomic component DNA-A of the 14 samples and one sequence of ∼580 bp corresponding to the AV1 gene was obtained from Cha157 sample.

PCR analysis from 14 RCA products performed with the degenerate primers PAC1C1960 and PAV1v722, which correspond to a partial DNA-A sequence. (A) (MW) contains a 100 bp DNA size marker, samples Po3696, Cha4265, Sq229 (Lane 1 –3), negative control (Lane 4) and positive control a bean sample infected by Bean golden yellow mosaic virus BGYMV (Lane 5) (B) (MW) contains a 500 bp DNA size marker, samples Sq142, Sq143, Sq150, Zu162, To166, Ch176, Es182, Sq185, Ch189, Ch201, Ch228, (Lane 1 –11), negative control (Lane 12) and positive control a bean sample infected by BGYMV (Lane 13).
The presence of SYMMoV or PepGMV was confirmed in those 15 samples. Seven sequences shared high similarity with SYMMoV (Table 1); and the remaining eight sequences were similar to PepGMV (Table 2). Only seven out of the latter eight sequences are shown in Table 2, because only the sequence corresponding to the capsid protein (AV1 gene) was obtained for sample Cha157. Nevertheless, the AV1 gene region is also useful for the preliminary identification of a virus species (Brown et al., 2001a). The nucleotide differences between these 15 isolates and other viral sequences obtained from the NCBI GenBank database were not clustered into particular regions of the DNA-A bottom or AV1 gene sequence.

Percent nucleotide identity among sequences of the DNA-A bottom of isolates of Squash yellow mild mottle virus (SYMMoV)

Percent nucleotide identity among sequences of the DNA-A bottom of isolates of Pepper golden mosaic virus (PepGMV)
The sequences similar to SYMMoV were from cucurbit samples and two were from solanaceous hosts (tomato and pepper). The sequence of the DNA-A bottom region (Table 1) shared 96 – 97% similarity with SYMMoV (GenBank, AY064391) and 89 – 90% with MCLCuV-GT (GenBank, AF325497). Four of the PepGMV sequences were from cucurbit samples, three were from pepper and one was from Erythrina spp. (Fabaceae). Sequences of the DNA-A bottom region (Table 2) shared 97 – 99% identity with PepGMV-CR (GenBank, AF149227) and 95 – 97% with PepGMV-Ser (GenBank, AY928516). The capsid protein gene sequence (AV1, ∼580 bp) from the chayote sample Cha157 shared 98% similarity with PepGMV-CR (GenBank, AF149227) and 95% PepGMV-YV (GenBank, AF063139).
Phylogenetic relationships among sequences corresponding to the DNA-A bottom region are shown in Fig. 2. Isolates from solanaceous plants (Ch189 and To166) and cucurbits (Sq142, Sq143, Sq185, Sq229 and Zu162) formed a well-defined group that was clearly related to the Costa Rican isolate of SYMMoV obtained by Karkashian et al. (2002). Isolates from cucurbits (Sq150, Es182, Cha4265), pepper (Ch176, Ch201, Ch228) and Erythrina sp. (Po3696) were grouped with isolates of PepGMV.
Discussion
Symptoms observed in the analyzed samples were similar to those previously described for plants infected with begomovirus (Hull, 2002) (Fig. 1). These viruses have caused negative effects on production in crops such as cotton, bean, tomato, pepper and cucurbits (Rojas et al., 2005). Moreover, begomoviruses are considered to be agronomically important pathogens in Costa Rica (Hilje et al., 1993) and worldwide (Morales, 2010; Navas-Castillo et al., 2011; Rojas and Gilbertson, 2008). The percent of nucleotide identity among partial sequences of the SYMMoV isolates detected in this study was between 97 and 100% (Table 1), indicating a low variability between them. Further more isolates and research are needed to confirm this observation. The phylogenetic relationships shown in Fig. 3 indicated that the partial sequence of the SYMMoV isolates obtained during this study are more closely related to MCLCuV-GT than to other begomoviruses infecting cucurbits. It has been considered that SYMMoV may be an isolate of MCLCuV (Brown et al., 2005). The sequence similarity for the DNA-A of the two viruses (88.8%) is at the limit 89% defined by Fauquet et al. (2008) to distinguish between isolates of the same species. However, there are differences in symptoms, hosts range and sequence similarity in conserved regions to suggest that further information is required to elucidate this case.

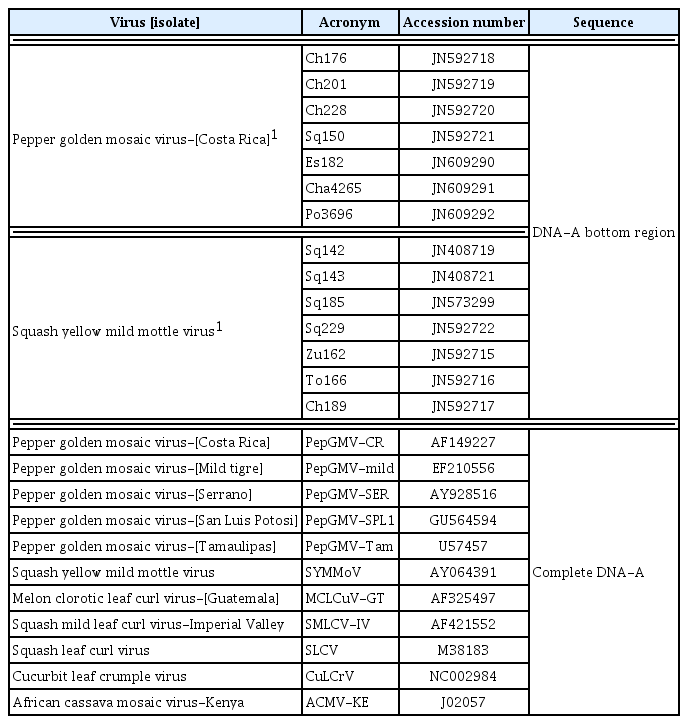
Phylogenetic relationships established by neighbor-joining analysis based on partial sequences of DNA-A (∼1,100 bp) of begomovirus species shown in Table 3. Horizontal line length is proportional to the number of nucleotide differences at each branch. Bootstrap values for 1,000 replicates were used. African cassava mosaic virus (ACMV-[KE]) was used as an outgroup.
The symptoms described for MCLCuV-GT and SYMMoV are different. MCLCuV-GT produced severe leaf deformation and reduced fruit set in Cucumis melo (melon) (Brown et al., 2001b). While, SYMMoV caused no symptoms in the fruit (J. Karkashian, personal communication) and leaf chlorotic mottle are often mild in Carica papaya (papaya) and in cucurbits (Karkashian et al., 2002). Additionaly, MCLCuV-GT and SYMMoV also differ in their hosts range. The first report of MCLCuV-GT was in melon (Brown et al., 2001b) and there have since been other report in this crop (Jord et al., 2005). MCLCuV-GT experimentally infects Cucurbita maxima, C. pepo, Citrullus lanatus, Phaseolus vulgaris (bean), Nicotiana benthamiana and Datura stramonium (Idris et al., 2008). On the other hand, natural infections of SYMMoV have been detected not only in cucurbits but also in papaya (Karkashian et al., 2002) and more recently in bean (Karkashian et al., 2011). This is an uncommon characteristic for begomoviruses, which tend to have a narrow host range (Rojas et al., 2005). Moreover, MCLCuV-GT has not been found infecting tomato or pepper even under experimental conditions (Idris et al., 2008), whereas SYMMoV was detected in both solanaceaous crops in this study.
Regardless the sequence similarity of 88.8% between SYMMoV and MCLCuV-GT complete DNA-A, their association is less evident when only one region as the capsid protein gene is considered. According to Brown et al. (2011) sequence similarity among the capsid protein genes and the common region of the genomic components of these two viruses was less than 87%. These differences were also observed with partial sequences of the capsid protein gene of two Nicaraguan isolates (GenBank, AJ8442150, AJ8442151) detected in Cucurbita argyrosperma (Ala-Poikela et al., 2005). Comparison of a ∼260 bp sequence from the Nicaraguan isolates (GenBank, AJ842150, AJ842151) and partial sequences of isolate Sq142 (Gen-Bank, JN408719) obtained in this study showed 94 – 98% nucleotide similarity with SYMMoV (GenBank, AY064391) and only 80 – 83% with MCLCuV-GT (GenBank, AF325497).
More biological data such as the ability to generate infectious reassortants, sap transmissibility and germplasm susceptibility; will help to clarify the relationship of MCLCuV-GT and SYMMoV differences. There are previous examples for parallel evolution of new crop-infecting begomoviruses species in multiple locations, including bean golden mosaic disease (Gilbertson et al., 1993) and African cassava mosaic (Varma and Malathi, 2003). These discrepancies in host range, symptoms and sequence difference in conserved regions, may indicate that these are two related but distinct virus species or that these are two isolates from the same species with divergent evolution (Brown et al., 2011).
The percent of nucleotide identity among partial sequences of PepGMV from this study was high (97 – 100%, Table 2), in concordance with a previous study with PepGMV isolates from North America (Brown et al., 2005). It should be noted that the partial sequence of sample Es182 showed lower similarity (97% and 95%) with the PepGMV-CR (GenBank, AF149227) and the PepGMV-Ser (GenBank, AY928516) than the other isolates of this study; however it is clearly an isolate of the same virus (Table 2). A close phylogenetic relationship (Fig. 3) was showed between samples in this study and the original Costa Rican PepGMV isolate obtained by Lotrakul et al. (2000) from Capsicum frutescens and C. chinense. There was also an association with other previously reported isolates of PepGMV from different geographic regions (Brown et al., 2005).
Diseases caused by PepGMV were previously reported in Costa Rica in pepper (Lotrakul et al., 2000) and tomato (Nakhla et al., 2005). Additionally, this virus has been detected in Cucurbita argyrosperma in Nicaragua (Ala-Poikela et al., 2005) as well as in cucurbits in this study. However, this is the first report of a begomovirus (PepGMV) in Sechium edule (chayote) in the Western Hemisphere. Prior to the current study, Chayote mosaic tymovirus (ChMV; Bernal et al., 2000; Hord et al., 1997) was the only known virus infecting chayote in Costa Rica. Although yield losses associated with PepGMV in chayote in the country are not reported yet; a more detailed study of the incidence and distribution of the virus in chayote plantations is required to establish the potential threat of this begomovirus. This is specially important when Costa Rica is one of the main countries producing and exporting chayote to the United States, Canada, Central America and the Caribbean (Cadena-Iñiguez et al., 2007).
Indigenous begomoviruses in other regions of the world apparently move between cucurbits and solanaceous crops. This is the case of Tomato leaf curl virus-New Delhi (ToLCV-NDe) in India (Mandal et al., 2004; Phaneendra et al., 2012; Sohrab et al., 2003; 2013), Squash leaf curl Philippines virus (SLCPHV) in Taiwan (Tsai et al., 2011) and Tomato leaf curl Palampur virus (ToLCPMV) reported in Iran (Heydarnejad et al., 2012). Probably PepGMV occurrence in chayote report in this study is also another example of this phenomenon. Additionally, the Chayote yellow mosaic virus (ChaYMV, GenBank AJ223191) detected in chayote plantations in the southeast of India and Nigeria, emerged as a potential new begomovirus specie.
The detection of PepGMV in bean (Hilje et al., 1993) and Erythrina spp. suggest the ability of this virus to infect also Fabaceae species. Although Erythrina spp. is not a crop for human consumption, it is used as a living fence (Muschler and Bonnemann, 1997) and in agroecosystems using a silvopastoral system (Morton, 1994). Close association of Erythrina spp. with whitefly infested vegetable crops may facilitate the transmission of the virus to this species when crops are eliminated. Erythrina trees may also serve as hosts for the virus in between cropping seasons and reservoirs for infection of new fields.
The results discussed herein may be an example of the adaptation of begomoviruses to new hosts and environments. Further studies are needed to confirm this hypothesis. The emergence of new viral strains and species due to the adaptation of existing viruses is a worldwide process that occurs constantly over time (Anderson et al., 2004; Fargette et al., 2006; Jones, 2009; Rojas and Gilbertson, 2008; Navas-Castillo et al., 2011). Therefore, it is recommended to perform periodical molecular characterization of begomoviruses, in order to determine the relationships between preexisting begomoviruses species and their hosts, and to identify arising new isolates/species. A clear understanding of the factors determining the evolution of these viruses is of importance in order to avoid future economic losses and to avoid their dispersal to other regions and crops.
Acknowledgements
The authors thank Felipe Arauz Cavallini and Mauricio Montero Astua for comments on the manuscript, and the National Council for Scientific and Technological Research (CONICIT), the Ministry of Science and Technology (MICIT), the State Fund for Higher Education (FEES) and Universidad de Costa Rica for financial support for this project.