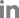|
 |
| Plant Pathol J > Volume 30(2); 2014 > Article |
Abstract
Lettuce mosaic virus (LMV) causes disease of plants in the family Asteraceae, especially lettuce crops. LMV isolates have previously been clustered in three main groups, LMV-Yar, LMV-Greek and LMVRoW. The first two groups, LMV-Yar and LMV-Greek, have similar characteristics such as no seed-borne transmission and non-resistance-breaking. The latter one, LMV-RoW, comprising a large percentage of the LMV isolates contains two large subgroups, LMV-Common and LMV-Most. To date, however, no Korean LMV isolate has been classified and characterized. In this study, LMV-Muju, the Korean LMV isolate, was isolated from lettuce showing pale green and mottle symptoms, and its complete genome sequence was determined. Classification method of LMV isolates based on nucleotide sequence divergence of the NIb-CP junction showed that LMV-Muju was categorized as LMV-Common. LMV-Muju was more similar to LMV-O (LMV-Common subgroup) than to LMV-E (LMV-RoW group but not LMV-Common subgroup) even in the amino acid domains of HC-Pro associated with pathogenicity, and in the CI and VPg regions related to ability to overcome resistance. Taken together, LMV-Muju belongs to the LMV-Common subgroup, and is expected to be a seed-borne, non-resistance-breaking isolate. According to our analysis, all other LMV isolates not previously assigned to a subgroup were also included in the LMV-RoW group.
Lettuce mosaic virus (LMV) belonging to the genus Potyvirus in the family Potyviridae is one of the most important pathogens of lettuce (Revers et al., 1997b). In most cases, LMV is seed-borne and transmitted by aphid in a non-persistent transmission manner (Bos et al., 1994; Nebreda et al., 2004). LMV has a single positive-stranded genomic RNA of 10,080 nucleotides in length and a poly A tail at its 3ŌĆ▓ end. The viral genomic RNA contains a single open reading frame (ORF) which encodes a single large polyprotein of 3,255 amino acids (Revers et al., 1997a). The genomic RNA of LMV is covalently linked to VPg, viral protein genome linked, at its 5ŌĆ▓ end and is encapsidated in capsid protein to form flexuous virions.
Worldwide, LMV is important disease of plants in the family Asteraceae, especially lettuce (German-Retana et al., 2008). Lettuce infected with LMV shows several symptoms such as mosaic, dwarfism, yellowing, wilting and failure to form heads. In order to protect lettuce crops, both genetic resistance and virus-free seed are generally used in the field. The recessive allelic resistance genes mo11 and mo12 correspond to alleles encoding the eukaryotic translation initiation factor elF4E (eIF4E) in lettuce crops (Tavert-Roudet et al., 2012). However, some LMV isolates have been overcoming the resistance induced by mo1 alleles in the last 20 years.
LMV isolates have previously been clustered in three main groups, LMV-Yar, LMV-Greek and LMV-RoW (Rest of the World) (German-Retana et al., 2008; Krause-Sakate et al., 2002). LMV-Yar has only one LMV isolate from Yemen and LMV-Greek has 10 LMV isolates from Greece. These two groups are non-seed-borne and non-resistance-breaking. Except for these 11 LMV isolates, all of the remaining isolates distributed around the globe are included in LMV-RoW containing two large subgroups, LMV-Common and LMV-Most (Krause-Sakate et al., 2004). Both subgroups are seed-borne, but only LMV-Most has the ability to overcome resistance triggered by recessive allelic resistance genes mo11 and mo12 (Bos et al., 1994; Pink et al., 1992; Revers et al., 1997a, b; Tavert-Roudet et al., 2012). Accordingly, LMV-Most was named after mo1-breaking and seed transmitted. Classification of LMV isolates is based on nucleotide sequence divergence of the nuclear inclusion protein b (NIb)-coat protein (CP) junction between positions 8,936 and 9,151 (Krause-Sakate et al., 2004).
So far, no Korean LMV isolate has been classified and characterized. In this study, we report the characteristics of LMV-Muju, Korean LMV isolate, infecting lettuce. In addition to LMV-Muju, we classified all the rest of the LMV isolates not included in previous report showing dendrogram of LMV isolates.
In order to analyze characteristics of the Korean LMV isolate, we collected lettuce showing pale green and mottle symptoms in emerging crops in Muju county, Jeollabukdo, Korea in August 2011 (Fig. 1a).
The reverse transcription (RT) polymerase chain reaction (PCR) assay was conducted for the detection of LMV. Total RNA was extracted from LMV-infected leaves with TRI Reagent (MRC, USA) according to the manufacturerŌĆÖs instructions. The forward primer 08894p (5ŌĆ▓- CCG TAC ATA GCI GAR TGT GCT -3ŌĆ▓) and the reverse primer 09171m (5ŌĆ▓- GCG TTG ATG TCG TCA TCY TT -3ŌĆ▓) giving product size of 278 bp were used as diagnostic PCR primers (Krause-Sakate et al., 2002; Peypelut et al., 2004). RT reaction using random N25 primer was performed with one cycle at 42┬░C for 1 h. PCR was performed with 32 cycles using the procedure (95┬░C, 30s; 50┬░C, 30s and 72┬░C, 1 min), followed by additional extension cycle at 72┬░C for 5 min. Amplified RT-PCR product was cloned into RBC TA Cloning Vector (RBC Bioscience, Taiwan) and sequence was determined by GenoTech Corp. sequencing service. We detected LMV from lettuce showing pale green and mottle symptoms by using RT-PCR. The sequence result showed sequence homology of up to 98% with characterized LMV isolates, so we therefore named the isolate LMV-Muju after its origin.
In order to prove pathogenicity of LMV-Muju, we inoculated test plants with extracts of infected leaf tissue based on KochŌĆÖs postulates. Seedlings of N. benthamiana were mechanically inoculated with crude sap of leaf sample infected with LMV-Muju in 1X phosphate buffered saline, pH 7.4, and grown in insect-free incubation room maintained at 23┬░C with 16 h light period. Disease symptoms of the test plants inoculated with LMV were observed at 10 days post-inoculation (dpi). At 21 dpi, each plant showed systemic mosaic symptoms (Fig. 1b). We also detected LMV from test plants by RT-PCR.
In order to determine the complete LMV genome sequence, 8 primer sets were designed from well conserved regions of four complete LMV sequences (GenBank accession no. X97704, X97705, AJ278854, and AJ306288) obtained from NCBI GenBank database (Table 1). Total RNA was extracted from LMV-Muju with the TRI Reagent (MRC, USA) and cDNA was synthesized using random N25 primer with RevertAid Reverse Transcriptase (Thermo scientific, USA) according to the manufacturersŌĆÖ instructions. PCR was performed with 32 cycles using the procedure (95┬░C, 30s; 53┬░C, 30s and 72┬░C, 2 min), followed by additional extension cycle at 72┬░C for 5 min. Amplified RT-PCR products were cloned and sequenced as above. Assembly of all the sequences was conducted using the DNAMAN version 5.0.10 program. The complete genome of LMV-Muju (GenBank accession no. KF955619) has 10,080 nt in length, including non-translated sequences at both termini, as for previously characterized isolates (Fig. 2).
Phylogenetic analysis was conducted using CLC Main Workbench 6.1.1 program. LMV-Muju was classified based on nucleotide sequence divergence of the NIb-CP junction between positions 8936 and 9151, and it was sorted into LMV-Common belonging to LMV-RoW (Fig. 2, 3). This result suggested that LMV-Muju may be seed-borne and a non-resistance-breaking isolate. Evidence from previous studies indicates that LMV isolates categorized in the same group share similar characteristics (German-Retana et al., 2008; Krause-Sakate et al., 2002). In addition to LMV-Muju, all the rest of the LMV isolates not included in previous studies were also classified (Fig. 3). As in previous studies, all of the remainder fall within the LMVRoW group except for the 11 LMV isolates in LMV-Yar and LMV-Greek.
The large single polyprotein of LMV is processed by three viral proteinases, P1, helper component proteinase (HC-Pro) and nuclear inclusion protein a (NIa)-Pro, into ten mature proteins playing roles in replication and survival of the virus in the host cell (Fig. 2). Several studies reported that HC-Pro proteins of the genus Potyvirus are symptom determinant(s) for pathogenicity (Atreya and Pirone, 1993; Atreya et al., 1992; Dolja et al., 1992; Klein et al., 1994). Ballut et al. (2005) reported that HC-Pro protein of LMV is a pathogenicity regulator which interferes with some of the catalytic functions of the 20S proteasome. LMV-O induces relatively mild mosaic, but LMV-E induces severe mosaic with malformation, stunting and/or necrosis (German-Retana et al., 2008; Pink et al., 1992; Redondo et al., 2001). Redondo et al. (2001) reported that HC-Pro protein of LMV-E is crucial for development of the severe symptoms in susceptible lettuce Trocad├®ro. Although the symptoms induced by LMV-0 and LMV-E are different, amino acid residues of HC-Pro proteins of the two isolates are quite similar (German-Retana et al., Revers et al., 1997a). They differ at only seven amino acid residues between positions 35 and 287 of HC-Pro proteins (Fig. 2). LMV-Muju was identical to LMV-O at these seven amino acid positions and in inducing relatively mild mosaic symptoms. There were only two different positions at amino acids 72 and 83 of LMV-Muju HC-Pro; however, LMV-O is identical to LMV-E at amino acids 72 and 83. These results suggested that LMV-Muju is similar to LMV-O in having the symptom determinants causing relatively mild mosaic as shown in Fig. 1a, and that the variant residues at positions 72 and 83 do not influence symptom development.
There are some LMV isolates having an ability to overcome resistance triggered by recessive allelic resistance mo1 gene (mo11 and mo12), encoding the eIF4E (Krause-Sakate et al., 2002; Revers et al., 1997a, b; Tavert-Roudet et al., 2012). Previous studies reported that the C terminus of the cylindrical inclusion (CI) protein and the VPg interacting with lettuce eIF4E are involved in mo1 resistance-breaking (Abdul-Razzak et al., 2009; Tavert-Roudet et al., 2012). Mutations in VPg allowed LMV-O to overcome mo11 resistance, but mutations in C-terminal region of CI protein allowed it to overcome both alleles (Abdul-Razzak et al., 2009). In other words, a mutation at amino acid residue 621 of the CI was sufficient to cause virulence on resistance lettuce (Fig. 2). LMV-E has threonine at amino acid 621 of the CI protein, but LMV-O and LMV-Muju have serine. This result is consistent with the classification shown schematically in Fig. 3. Taken all together, LMV-Muju belonging to LMV-Common is expected to be a seed-borne and non-resistance-breaking isolate.
Acknowledgments
This study was supported by a grant (Project No. 609002-5) from the Screening Center for Disease Resistant Vegetable Crops of Technology Development Program for Agriculture and Forestry, Ministry for Food, Agriculture, Forestry and Fisheries, Republic of Korea, and carried out with the support of ŌĆ£Cooperative Research Program for Agricultural Science & Technology Development (Project No. PJ00912703)ŌĆØ, Rural Development Administration, Republic of Korea.
Fig.┬Ā1.
Disease symptoms induced by LMV. (A) Systemic mild mosaic symptoms of lettuce naturally infected with LMV. (B) N. benthamiana test plant was inoculated with sap of LMV-infected lettuce and showed systemic mild mosaic symptoms.

Fig.┬Ā2.
Schematic representation of the complete LMV genome. LMV isolates are classified based on nucleotide sequence divergence of the NIb-CP junction between positions 8,936 and 9,151. HC-Pro protein of LMV is regarded as a pathogenicity regulator. Amino acid residue 621 of the CI of LMV is involved in overcoming resistance triggered by mo1 alleles.
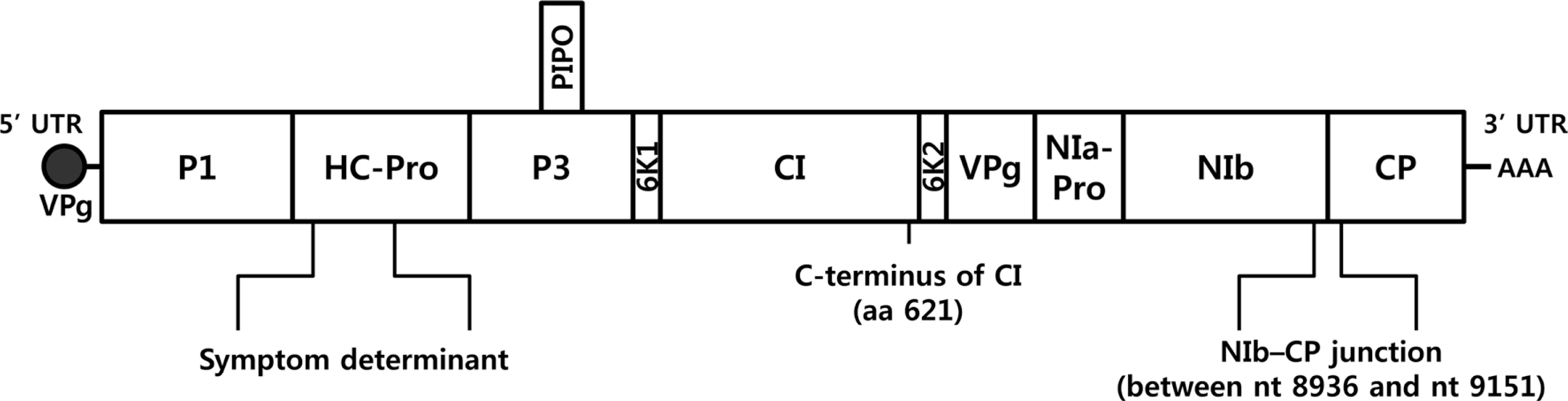
Fig.┬Ā3.
Phylogenetic tree showing nucleotide sequence divergence of the NIb-CP junction of LMV isolates (nt 8,936-9,151). The length of the branches and percentage of bootstrap frequency are indicated. LMV-Muju was categorized to the LMV-Common subgroup within the LMV-RoW group, and all the rest of the LMV isolates not included in previous reports were also sorted into LMV-RoW.
Table┬Ā1.
The primer sets used for complete genome of LMV-Muju in RT-PCR
References
Abdul-Razzak, A, Guiraud, T, Peypelut, M, Walter, J, Houvenaghel, MC, Candresse, T, Gall, OL and German-Retana, S 2009. Involvement of the cylindrical inclusion (CI) protein in the overcoming of an eIF4E-mediated resistance against Lettuce mosaic potyvirus. Mol Plant Pathol. 10:109-113.



Atreya, CD, Atreya, PL, Thornbury, DW and Pirone, TP 1992. Site-directed mutations in the potyvirus HC-Pro gene affect helper component activity, virus accumulation, and symptom expression in infected tobacco plants. Virology. 191:106-111.


Atreya, CD and Pirone, TP 1993. Mutational analysis of the helper component-proteinase gene of a potyvirus: effects of amino acid substitutions, deletions, and gene replacement on virulence and aphid transmissibility. Proc Natl Acad Sci USA. 90:11919-11923.



Ballut, L, Drucker, M, Pugni├©re, M, Cambon, F, Blanc, S, Roquet, F, Candresse, T, Schmid, HP, Nicolas, P, Gall, OL and Badaoui, S 2005. HcPro, a multifunctional protein encoded by a plant RNA virus, targets the 20S proteasome and affects its enzymic activities. J Gen Virol. 86:2595-2603.


Bos, L, Huijberts, N and Cuperus, C 1994. Further observations on variation of lettuce mosaic virus in relation to lettuce (Lactuca sativa), and a discussion of resistance terminology. Eur J Plant Pathol. 100:293-314.


Dolja, VV, McBride, HJ and Carrington, JC 1992. Tagging of plant potyvirus replication and movement by insertion of beta-glucuronidase into the viral polyprotein. Proc Natl Acad Sci USA. 89:10208-10212.



German-Retana, S, Walter, J and Gall, OL 2008. Lettuce mosaic virus: from pathogen diversity to host interactors. Mol Plant Pathol. 9:127-136.


Klein, PG, Klein, RR, Rodr├Łguez-Cerezo, E, Hunt, AG and Shaw, JG 1994. Mutational analysis of the tobacco vein mottling virus genome. Virology. 204:759-769.


Krause-Sakate, R, Gall, OL, Fakhfakh, H, Peypelut, M, Marrakchi, M, Varveri, C, Pavan, MA, Souche, S, Lot, H, Zerbini, FM and Candresse, T 2002. Molecular and Biological characterization of Lettuce mosaic virus (LMV) isolates reveals a distinct and widespread type of resistance-breaking isolate: LMV-Most. Phytopathology. 92:563-572.


Krause-Sakate, R, Fakhfakh, H, Peypelut, M, Pavan, MA, Zerbini, FM, Marrakchi, M, Candresse, T and Gall, OL 2004. A naturally occurring recombinant isolate of Lettuce mosaic virus. Arch Virol. 149:191-197.



Nebreda, M, Moreno, A, P├®rez, N, Palacios, I, Seco-Fern├Īndez, V and Fereres, A 2004. Activity of aphids associated with lettuce and broccoli in Spain and their efficiency as vectors of Lettuce mosaic virus. Virus Res. 100:83-88.


Peypelut, M, Krause-Sakate, R, Guiraud, T, Pavan, MA, Candresse, T, Zerbini, FM and Gall, OL 2004. Specific detection of lettuce mosaic virus isolates belonging to the ŌĆ£MostŌĆØ type. J. Virol. Methods. 121:119-124.


Pink, DAC, Lot, H and Johnson, R 1992. Novel pathotypes of lettuce mosaic virus - breakdown of a durable resistance? Euphytica. 63:169-174.


Redondo, E, Krause-Sakate, R, Yang, SJ, Lot, H, Gall, OL and Candresse, T 2001. Lettuce mosaic virus pathogenicity determinants in susceptible and tolerant lettuce cultivars map to different regions of the viral genome. Mol Plant-Microbe Interact. 14:804-810.


Revers, F, Yang, SJ, Walter, J, Souche, S, Lot, H, Gall, OL, Candresse, T and Dunez, J 1997a. Comparison of the complete nucleotide sequences of two isolates of lettuce mosaic virus differing in their biological properties. Virus Res. 47:167-177.


Revers, F, Lot, H, Souche, S, Gall, OL, Candresse, T and Dunez, J 1997b. Biological and molecular variability of lettuce mosaic virus isolates. Phytopathology. 87:397-403.


Tavert-Roudet, G, Abdul-Razzak, A, Doublet, B, Walter, J, Delaunay, T, German-Retana, S, Michon, T, Gall, OL and Candresse, T 2012. The C terminus of lettuce mosaic potyvirus cylindrical inclusion helicase interacts with the viral VPg and with lettuce translation eukaryotic initiation factor 4E. J Gen Virol. 93:184-193.


- TOOLS
-
METRICS

- Related articles
-
Characteristics of Cucumber mosaic virus Infecting Zucchini in Korea2010 June;26(2)
Isolation and Characterization of Pepper mottle virus Infecting Tomato in Korea2008 June;24(2)
Characterization of Colletotrichum Isolates Causing Anthracnose of Pepper in Korea2008 March;24(1)



 PDF Links
PDF Links PubReader
PubReader Full text via DOI
Full text via DOI Full text via PMC
Full text via PMC Download Citation
Download Citation Print
Print